The JAK-STAT Pathway in Neuroinflammation: Molecular Mechanisms, Therapeutic Targeting, and Recent Advances
This comprehensive review details the molecular mechanisms of JAK-STAT pathway activation in neuroinflammatory contexts, a critical signaling axis in neurological disorders like multiple sclerosis, Alzheimer's disease, and stroke.
The JAK-STAT Pathway in Neuroinflammation: Molecular Mechanisms, Therapeutic Targeting, and Recent Advances
Abstract
This comprehensive review details the molecular mechanisms of JAK-STAT pathway activation in neuroinflammatory contexts, a critical signaling axis in neurological disorders like multiple sclerosis, Alzheimer's disease, and stroke. It explores the foundational biology, from cytokine-receptor binding to nuclear translocation and gene regulation. Methodological approaches for studying pathway dynamics in neural and glial cells are examined, alongside current and emerging therapeutic strategies using JAK inhibitors (JAKi). The article addresses common experimental challenges, data interpretation pitfalls, and optimization techniques for in vitro and in vivo models. Finally, it provides a comparative analysis of JAK-STAT's role across different neuroinflammatory diseases and validates its therapeutic relevance through clinical and preclinical evidence, offering a roadmap for researchers and drug development professionals targeting this pathway for neurological therapeutics.
Decoding the Signal: Core Mechanisms of JAK-STAT Activation in the Inflamed CNS
Within the broader context of elucidating the JAK-STAT pathway's mechanism of activation in neuroinflammation, this guide details the fundamental cellular and molecular players driving neuroinflammatory responses. Neuroinflammation, a complex process central to numerous neurological disorders, is characterized by the activation of glial cells and the release of soluble mediators, many of which signal through the JAK-STAT cascade.
Core Cellular Players in Neuroinflammation
Resident CNS Cells
- Microglia: The primary innate immune effector cells of the CNS. In response to insult, they shift from a surveillant (M2-like) to an activated (M1-like) state, releasing pro-inflammatory cytokines and reactive oxygen species.
- Astrocytes: Provide metabolic and structural support. During neuroinflammation, they become reactive (astrogliosis), contributing to both pro- and anti-inflammatory signaling, and can disrupt blood-brain barrier (BBB) integrity.
- Oligodendrocytes: Myelin-producing cells vulnerable to inflammatory damage, linking neuroinflammation to demyelination.
- Endothelial Cells: Form the BBB; their activation facilitates leukocyte infiltration.
Infiltrating Peripheral Immune Cells
Following BBB compromise, peripheral cells infiltrate the CNS parenchyma.
- Monocyte-derived Macrophages: Augment the pro-inflammatory milieu.
- Lymphocytes (T-cells, B-cells): Drive adaptive immune responses. T helper 1 (Th1) and Th17 cells are particularly implicated in propagating inflammation.
Key Cytokines and Their Signaling Pathways
Cytokines are the primary communicators between these cells. A critical subset activates the JAK-STAT pathway, a central thesis focus.
Pro-inflammatory Cytokines (Drivers)
These cytokines are potent activators of microglia and astrocytes, and are major upstream activators of the JAK-STAT pathway.
Table 1: Key Pro-inflammatory Cytokines in Neuroinflammation
| Cytokine | Primary Cellular Source | Primary JAK/STAT Pathway Engaged | Key Functions in Neuroinflammation |
|---|---|---|---|
| IL-1β | Microglia, Macrophages | Indirect modulation | Pyrogen, promotes BBB breakdown, enhances astrocyte reactivity, induces other cytokines (e.g., IL-6). |
| TNF-α | Microglia, Astrocytes, T-cells | JAK1/2 - STAT1/3/5 (non-canonical) | Induces apoptosis, activates microglia, disrupts BBB, synergizes with IFN-γ. |
| IL-6 | Microglia, Astrocytes, Endothelial cells | JAK1/2 - STAT3 (canonical) | Acute phase response, B/T-cell differentiation, driver of astrogliosis, key for Th17 differentiation. |
| IFN-γ | Infiltrating T-cells, NK cells | JAK1/2 - STAT1 (canonical) | Potent microglial activator, promotes antigen presentation, upregulates MHC molecules. |
| IL-12/IL-23 | Microglia, Macrophages, Dendritic cells | JAK2/TYK2 - STAT4 (IL-12), STAT3 (IL-23) | Polarize T-cells toward Th1 (IL-12) or stabilize Th17 (IL-23) phenotypes. |
Anti-inflammatory & Resolution Cytokines (Modulators)
These cytokines often signal via JAK-STAT to counterbalance inflammation and promote repair.
Table 2: Key Anti-inflammatory & Resolution Cytokines
| Cytokine | Primary Cellular Source | Primary JAK/STAT Pathway Engaged | Key Functions in Neuroinflammation |
|---|---|---|---|
| IL-10 | Microglia (M2), Tregs, Astrocytes | JAK1/TYK2 - STAT3 | Decreases pro-inflammatory cytokine production, promotes M2 microglial phenotype. |
| TGF-β | Microglia, Astrocytes, Tregs | SMAD pathway (primary) | Suppresses microglial activation, promotes regulatory T-cell functions, involved in glial scar formation. |
| IL-4 / IL-13 | T-cells, Mast cells | JAK1/3 - STAT6 | Promote alternative (M2) microglial/macrophage activation, tissue repair, and remyelination. |
The Central Axis: JAK-STAT Pathway Activation
The JAK-STAT pathway is the principal signaling mechanism for many neuroinflammatory cytokines. Its activation is a multi-step process:
- Cytokine Binding: A cytokine binds to its cognate transmembrane receptor, inducing dimerization or conformational change.
- JAK Activation: Receptor-associated Janus Kinases (JAKs) trans-phosphorylate each other, becoming fully activated.
- STAT Recruitment & Phosphorylation: STAT transcription factors are recruited to receptor phospho-tyrosine motifs and are phosphorylated by JAKs.
- STAT Dimerization & Nuclear Translocation: Phosphorylated STATs dimerize, translocate to the nucleus, and bind specific DNA sequences to regulate gene transcription.
Experimental Protocols for Key Analyses
Protocol: Assessing Microglial ActivationIn Vitro
Aim: To stimulate and characterize the inflammatory phenotype of the BV2 microglial cell line or primary microglia.
- Cell Culture: Maintain BV2 cells in DMEM + 10% FBS. For activation, seed cells at 2.5 x 10^5 cells/mL in 6-well plates.
- Stimulation: Treat cells with a cytokine cocktail (e.g., 100 ng/mL LPS + 20 ng/mL IFN-γ) for 6-24 hours. Include vehicle control.
- RNA/Protein Harvest: Harvest cells for qPCR (inflammatory markers: Tnf, Il1b, Nos2, Cd86) or western blot (iNOS, phospho-STAT1/3).
- Supernatant Analysis: Collect supernatant for ELISA quantification of secreted TNF-α, IL-6.
Protocol: Co-culture to Study Neuron-Glia Interactions
Aim: To model neuroinflammatory toxicity.
- Neuron Culture: Plate primary cortical neurons in transwell inserts (pore size 0.4 µm) at 1 x 10^5 cells/insert. Culture for 7-10 days in vitro (DIV).
- Glial Culture/Stimulation: Plate microglia in the bottom well. At experiment day, stimulate microglia with inflammatory agents (e.g., 100 ng/mL LPS).
- Co-culture: Place neuron-containing insert into the well with stimulated microglia. Co-culture for 24-72 hours.
- Assessment: Analyze neurons in the insert for viability (MTT assay), synaptic density (immunofluorescence for PSD-95, Synapsin), or apoptosis (caspase-3 cleavage).
Protocol: Phospho-STAT Analysis via Western Blot
Aim: To detect activation of the JAK-STAT pathway in brain tissue or cell lysates.
- Lysis: Homogenize tissue or cells in RIPA buffer supplemented with phosphatase and protease inhibitors.
- Electrophoresis: Load 20-30 µg of protein per lane on a 8-10% SDS-PAGE gel.
- Transfer & Blocking: Transfer to PVDF membrane, block with 5% BSA in TBST for 1 hour.
- Antibody Incubation: Incubate with primary antibodies overnight at 4°C:
- Phospho-specific: anti-pSTAT1 (Tyr701), anti-pSTAT3 (Tyr705).
- Total protein: anti-STAT1, anti-STAT3.
- Loading control: anti-β-actin.
- Detection: Use HRP-conjugated secondary antibodies and chemiluminescent substrate. Quantify band density.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Neuroinflammation & JAK-STAT Research
| Reagent Category | Specific Example(s) | Function & Application |
|---|---|---|
| Recombinant Cytokines | Mouse/rHu IFN-γ, IL-6, IL-1β, IL-4, LPS (TLR4 agonist) | Used to stimulate glial cells in vitro or in vivo to induce inflammatory responses and activate specific pathways (e.g., JAK-STAT). |
| JAK-STAT Inhibitors | Ruxolitinib (JAK1/2 inhibitor), Tofacitinib (JAK1/3 inhibitor), STAT3 Inhibitor VI (S3I-201) | Pharmacological tools to dissect pathway contribution to neuroinflammatory phenotypes in vitro and in disease models. |
| Phospho-Specific Antibodies | Anti-phospho-STAT1 (Tyr701), Anti-phospho-STAT3 (Tyr705) | Critical for detecting pathway activation via western blot, immunohistochemistry, or flow cytometry. |
| ELISA Kits | Mouse TNF-α, IL-6, IL-1β Quantikine ELISA | Gold-standard for quantitative measurement of cytokine levels in cell culture supernatant, CSF, or brain homogenates. |
| Microglial Markers | Anti-IBA1 (ionized calcium-binding adapter molecule 1), Anti-TMEM119 | Immunohistochemical identification and quantification of microglia in tissue sections. IBA1 labels all microglia/macrophages; TMEM119 is more specific for resting microglia. |
| Flow Cytometry Antibodies | Anti-CD11b (Microglia/Macrophages), Anti-CD45 (Leukocytes), Anti-Ly6C (Monocyte subsets) | Used for immunophenotyping of CNS immune cells isolated from brain tissue via Percoll gradient, allowing differentiation of resident microglia from infiltrating macrophages. |
Understanding the interplay between key cytokines (notably IL-6, IFN-γ, IL-1β) and cellular players (microglia, astrocytes, infiltrating lymphocytes) provides the foundational context for investigating the JAK-STAT pathway's specific role. This pathway serves as a critical signaling nexus, translating extracellular inflammatory signals into sustained changes in gene expression within the CNS. Current drug development focuses heavily on targeting this axis, with JAK inhibitors being evaluated for their potential to modulate detrimental neuroinflammation while preserving protective functions. Future research must continue to delineate the spatiotemporal activation patterns of specific JAK-STAT modules in different cell types to enable precise therapeutic intervention.
1. Introduction & Thesis Context The Janus Kinase-Signal Transducer and Activator of Transcription (JAK-STAT) pathway is a principal signaling cascade transmitting extracellular cytokine signals directly to the nucleus, governing gene expression programs critical in immunity, proliferation, and differentiation. Its dysregulation is a cornerstone of numerous pathologies, including neuroinflammatory diseases. Within the context of neuroinflammation research, understanding the canonical JAK-STAT mechanism is paramount, as its hyperactivation in microglia, astrocytes, and infiltrating immune cells drives the production of pro-inflammatory mediators, contributing to neurodegeneration in conditions like multiple sclerosis, Alzheimer's disease, and stroke. This whitepaper details the core components, regulatory families, and experimental methodologies essential for investigating this pathway in neurological contexts.
2. Canonical Pathway Structure & Mechanism The canonical pathway is initiated by the binding of cytokines (e.g., IFN-γ, IL-6 family) to their cognate type I or II transmembrane receptors, which are constitutively associated with JAKs.
- Step 1: Ligand-induced receptor dimerization brings paired JAKs into proximity, leading to their trans-phosphorylation and activation.
- Step 2: Activated JAKs phosphorylate specific tyrosine residues on the receptor cytoplasmic tails, creating docking sites for STAT proteins via their Src Homology 2 (SH2) domains.
- Step 3: STATs are recruited and subsequently phosphorylated by JAKs on a conserved C-terminal tyrosine residue.
- Step 4: Phosphorylated STATs dissociate from the receptor, form homo- or heterodimers via reciprocal phospho-tyrosine-SH2 domain interactions.
- Step 5: STAT dimers translocate to the nucleus, bind specific gamma-activated sequence (GAS) elements in target gene promoters, and regulate transcription.
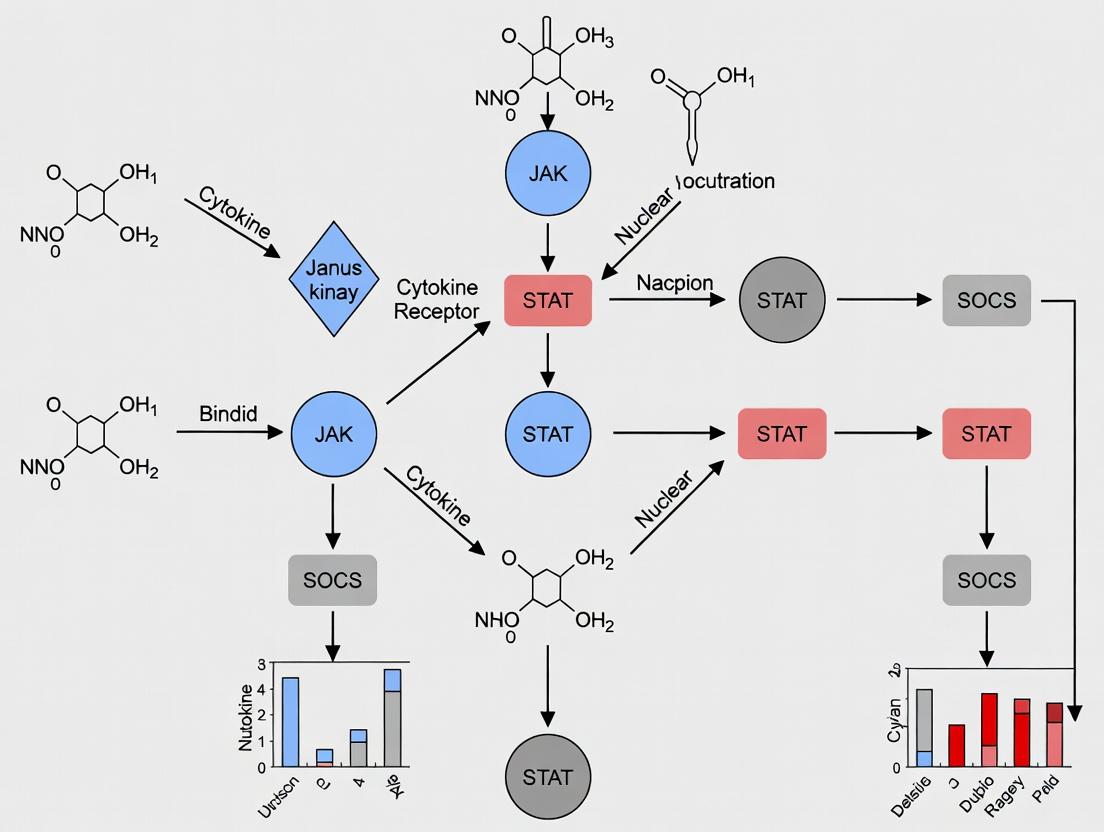
Diagram: Canonical JAK-STAT Activation Cascade
3. Core Component Families
3.1 Janus Kinases (JAKs) JAKs are non-receptor tyrosine kinases. Mammals express four JAKs: JAK1, JAK2, JAK3, and TYK2. Each pairs with specific cytokine receptor subunits.
Table 1: JAK Family Members, Association, and Key Functions
| JAK | Chromosome (Human) | Size (aa) | Primary Receptor Associations | Key Cytokine Signals | Phenotype of KO/Mutation |
|---|---|---|---|---|---|
| JAK1 | 1p31.3 | 1154 | GP130, IFNAR, IFNGR, γc-chain | IFN-α/β/γ, IL-6 family, IL-2, IL-4 | Perinatal lethality, neurologic deficits, defective IFN response. |
| JAK2 | 9p24.1 | 1132 | EPOR, TPOR, G-CSFR, GP130 | EPO, TPO, GH, IL-3, IL-5, IL-6 family | Embryonic lethality (E12.5) due to defective erythropoiesis. |
| JAK3 | 19p13.11 | 1124 | γc-chain | IL-2, IL-4, IL-7, IL-9, IL-15, IL-21 | Severe Combined Immunodeficiency (SCID). |
| TYK2 | 19p13.2 | 1187 | IFNAR, IL-12R, IL-23R | IFN-α/β, IL-12, IL-23 | Hyper-susceptibility to viral/bacterial infection, mild SCID. |
3.2 Signal Transducers and Activators of Transcription (STATs) Seven mammalian STATs (STAT1, STAT2, STAT3, STAT4, STAT5A, STAT5B, STAT6) translate phosphorylation into transcriptional programs.
Table 2: STAT Family Members, Key Activation Signals, and Functions
| STAT | Size (aa) | Primary Activating Cytokines | Key Target Genes | Major Biological Role |
|---|---|---|---|---|
| STAT1 | 750 | IFN-α/β/γ, IL-2, IL-6 | IRF1, CASP1, SOCS1 | Antiviral response, MHC class II upregulation, Th1 differentiation. |
| STAT2 | 851 | IFN-α/β | ISGF3 complex (with STAT1:IRF9) | Type I IFN antiviral response. |
| STAT3 | 770 | IL-6 family, IL-10, EGF, LIF | SOCS3, BCL2, MYC, GFAP | Acute phase response, glial activation, cell survival, oncogenesis. |
| STAT4 | 748 | IL-12, IFN-α | IFN-γ, IL-18R | Th1 differentiation, cell-mediated immunity. |
| STAT5A/B | 794/787 | IL-2, IL-3, IL-5, IL-7, GH, EPO | BCL2, CYCLIN D1, CIS | Lymphocyte proliferation, milk protein expression, erythropoiesis. |
| STAT6 | 847 | IL-4, IL-13 | CD23, MHC Class II, ARG1 | Th2 differentiation, B-cell class switching to IgE. |
4. Negative Regulatory Families
4.1 Suppressors of Cytokine Signaling (SOCS) SOCS proteins (CIS, SOCS1-7) are inducible feedback inhibitors. They function via: 1) Competitive binding to phospho-tyrosine sites on receptors/JAKs (SH2 domain), 2) Direct inhibition of JAK kinase activity (KIR domain in SOCS1/3), 3) Targeting bound proteins for proteasomal degradation via a SOCS-box E3 ubiquitin ligase complex.
4.2 Protein Inhibitors of Activated STATs (PIAS) PIAS proteins (PIAS1, PIAS3, PIASx, PIASy) regulate STAT signaling in the nucleus via multiple mechanisms: 1) Blocking STAT-DNA binding, 2) Promoting SUMOylation of STATs and other transcription factors, 3) Recruiting transcriptional co-repressors, 4) Modulating chromatin structure.
Diagram: JAK-STAT Negative Feedback Regulation
5. Experimental Protocols for Neuroinflammation Research
5.1 Assessing STAT Phosphorylation (Activation) in Glial Cells
- Objective: Determine temporal dynamics of STAT1/3 phosphorylation in primary microglia stimulated with IFN-γ or IL-6.
- Protocol:
- Cell Culture & Stimulation: Isolate primary microglia from postnatal rodent brains. Serum-starve for 4h. Stimulate with IFN-γ (50 ng/mL) or IL-6 (50 ng/mL + soluble IL-6Rα (50 ng/mL)) for 0, 15, 30, 60, 120 minutes.
- Cell Lysis: Lyse cells in RIPA buffer supplemented with phosphatase and protease inhibitors.
- Immunoblotting: Resolve 20-30 µg protein by SDS-PAGE. Transfer to PVDF membrane.
- Detection: Probe with primary antibodies: anti-pSTAT1 (Tyr701), anti-total STAT1, anti-pSTAT3 (Tyr705), anti-total STAT3. Use HRP-conjugated secondary antibodies and chemiluminescent substrate.
- Analysis: Quantify band density; express pSTAT levels normalized to total STAT.
5.2 Nuclear Translocation Assay via Immunofluorescence
- Objective: Visualize STAT3 nuclear translocation in astrocytes.
- Protocol:
- Culture & Stimulation: Plate astrocytes on poly-D-lysine-coated coverslips. Stimulate with IL-6 family cytokine (e.g., CNTF, 50 ng/mL) for 30 min.
- Fixation & Permeabilization: Fix with 4% PFA for 15 min. Permeabilize with 0.2% Triton X-100 for 10 min.
- Staining: Block with 5% BSA. Incubate with anti-STAT3 antibody (1:250) overnight at 4°C, followed by Alexa Fluor 488-conjugated secondary antibody (1:500) and DAPI (1 µg/mL) for 1h.
- Imaging: Acquire images using a confocal microscope. Analyze co-localization of STAT3 signal (green) with DAPI (blue) in the nucleus.
5.3 SOCS3 Feedback Induction Analysis (qRT-PCR)
- Objective: Measure SOCS3 mRNA induction as a pathway feedback readout.
- Protocol:
- Stimulation & RNA Extraction: Stimulate BV-2 microglial cells with IL-6 (as above) for 0, 30, 60, 120 min. Extract total RNA using TRIzol.
- cDNA Synthesis: Perform reverse transcription with 1 µg RNA using oligo(dT) primers.
- Quantitative PCR: Prepare SYBR Green master mix. Use primers:
- SOCS3-F: 5'-CACCTCTGACGAGACCAAACG-3'
- SOCS3-R: 5'-GTCACTGCGCTCCAGTAGAA-3'
- GAPDH-F: 5'-AGGTCGGTGTGAACGGATTTG-3'
- GAPDH-R: 5'-TGTAGACCATGTAGTTGAGGTCA-3'
- Analysis: Calculate ΔΔCt values; express SOCS3 mRNA levels relative to unstimulated control, normalized to GAPDH.
6. The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for JAK-STAT Neuroinflammation Research
| Reagent Category | Specific Example(s) | Function/Application |
|---|---|---|
| Recombinant Cytokines | Mouse/Rat IFN-γ, IL-6, IL-4, IL-10, CNTF, LIF. | Pathway activation in cell culture or in vivo models. |
| JAK Inhibitors (Tool Compounds) | Pyridine 6 (JAK1/2/3 inhibitor), Tofacitinib (JAK1/3 inhibitor), Ruxolitinib (JAK1/2 inhibitor). | Pharmacological inhibition to probe pathway function. |
| Phospho-Specific Antibodies | Anti-pSTAT1 (Tyr701), Anti-pSTAT3 (Tyr705), Anti-pJAK2 (Tyr1007/1008). | Detection of pathway activation via WB, IHC, or flow cytometry. |
| Total Protein Antibodies | Anti-STAT1, STAT3, STAT6, JAK1, JAK2, TYK2. | Loading controls and expression level assessment. |
| SOCS/PIAS Antibodies | Anti-SOCS1, SOCS3, PIAS1, PIAS3. | Studying negative regulator expression and localization. |
| ELISA/Kits | Phospho-STAT1/3/5 (Multi-pathway) Cellular Assay Kits; Mouse IFN-γ ELISA Kit. | Quantifying activation or cytokine levels in samples. |
| siRNA/shRNA | Pre-designed siRNA pools against human/mouse JAK1, STAT3, SOCS3. | Gene knockdown for functional studies. |
| Reporter Constructs | pSTAT3-TA-luc (STAT3-responsive luciferase), pGAS-luc. | Measuring STAT transcriptional activity in cell-based assays. |
The JAK-STAT signaling pathway is a principal mediator of cytokine and growth factor signaling, playing a central role in immune regulation and neuroinflammatory processes. In neuroinflammation, aberrant activation of this pathway by cytokines like IL-6, IFN-γ, and IL-1β drives glial cell activation, leukocyte infiltration, and neuronal damage, contributing to the pathogenesis of conditions such as multiple sclerosis, Alzheimer's disease, and neuropathic pain. The initial, critical triggering event is the engagement of specific cell-surface cytokine receptors, leading to the transphosphorylation and activation of receptor-associated Janus Kinases (JAKs). This molecular event serves as the essential "on-switch" for the entire downstream cascade, making its detailed understanding a priority for therapeutic intervention.
Core Molecular Machinery
Cytokine Receptors and JAK Association
Cytokine receptors involved in neuroinflammation lack intrinsic kinase activity. They instead rely on constitutively associated JAK family members (JAK1, JAK2, JAK3, TYK2). Receptor families are defined by their structural motifs and associated JAKs.
Table 1: Key Cytokine Receptor Complexes in Neuroinflammation
| Receptor Complex | Cytokine Ligands (Examples) | Associated JAKs | Primary CNS Cell Types | Role in Neuroinflammation |
|---|---|---|---|---|
| gp130 Family | IL-6, CNTF, LIF | JAK1, JAK2, TYK2 | Astrocytes, Microglia, Neurons | Astrogliosis, Acute phase response, Neural survival/damage |
| IFN-γ Receptor | IFN-γ | JAK1, JAK2 | Microglia, Astrocytes, Endothelial cells | MHC class II upregulation, Microglial activation, Blood-brain barrier disruption |
| Common Gamma (γc) Chain | IL-2, IL-4, IL-7 (limited CNS) | JAK1, JAK3 | Infiltrating T-cells, Microglia | T-cell proliferation & survival, Alternative glial activation |
| IFN-α/β Receptor | Type I IFNs | JAK1, TYK2 | All neural cells | Antiviral response, Modulator of MS pathology |
The Transphosphorylation Event
Upon ligand-induced receptor dimerization/oligomerization, the associated JAKs are brought into close proximity. This allows one JAK to phosphorylate a key tyrosine residue (Y1038/Y1039 in JAK2 kinase domain) on its partner JAK. This trans-phosphorylation event stabilizes the active conformation of the JAK kinase domain, dramatically increasing its catalytic activity. Activated JAKs then phosphorylate specific tyrosine residues on the intracellular receptor tails, creating docking sites for STAT proteins.
Experimental Methodologies for Analysis
Protocol: Co-Immunoprecipitation (Co-IP) and Western Blot to Detect JAK-Receptor Association and Transphosphorylation
Objective: To validate the physical interaction between a cytokine receptor and its associated JAK, and to detect ligand-induced JAK transphosphorylation.
Materials:
- Cell line (e.g., primary murine microglia, human astrocytoma cell line U87)
- Recombinant cytokine (e.g., IFN-γ, IL-6)
- Cell lysis buffer (RIPA buffer supplemented with phosphatase and protease inhibitors)
- Antibodies: Anti-receptor antibody (for IP), anti-JAK antibody, anti-phospho-JAK (e.g., p-JAK2 (Tyr1007/1008)), anti-STAT1, anti-p-STAT1 (Tyr701)
- Protein A/G magnetic beads
- SDS-PAGE and Western blotting system
Procedure:
- Stimulation: Serum-starve cells for 4-6 hours. Treat experimental groups with cytokine (e.g., 50 ng/mL IFN-γ for 15 min). Maintain an unstimulated control.
- Lysis: Rapidly wash cells with ice-cold PBS and lyse in IP lysis buffer. Clear lysates by centrifugation (14,000 x g, 15 min, 4°C).
- Immunoprecipitation: Incubate 500 µg of total protein with 2-5 µg of anti-receptor antibody overnight at 4°C. Add protein A/G beads for 2 hours. Wash beads stringently 3-4 times with lysis buffer.
- Elution: Elute bound proteins by boiling beads in 2X Laemmli sample buffer.
- Western Blot: Resolve proteins by SDS-PAGE, transfer to PVDF membrane, and probe sequentially.
- First, probe with anti-p-JAK and anti-JAK to assess JAK activation in the receptor complex.
- Strip membrane and re-probe with anti-receptor antibody to confirm successful IP.
- Downstream Validation: Analyze whole-cell lysates (non-IP) for total and phosphorylated STAT proteins to confirm functional pathway activation.
Protocol: In Vitro Kinase Assay for JAK Activity
Objective: To directly measure the enzymatic activity of JAKs immunoprecipitated from stimulated cells.
Materials:
- JAK immunoprecipitates (from protocol 3.1, but using anti-JAK antibody for IP)
- Kinase reaction buffer (HEPES, MgCl₂, MnCl₂, DTT)
- ATP (with [γ-³²P]ATP for radiometric assay or cold ATP for phospho-specific antibody detection)
- Recombinant substrate (e.g., GST-STAT1 or a poly-peptide corresponding to the receptor tail)
- Phosphocellulose paper (for radiometric assay) or SDS-PAGE equipment
Procedure:
- Prepare IP Complex: Perform IP as in 3.1 using an anti-JAK antibody. Wash beads twice with kinase wash buffer, then once with kinase reaction buffer.
- Kinase Reaction: Resuspend beads in 30 µL kinase reaction buffer containing 10 µM ATP (and 5 µCi [γ-³²P]ATP if radioactive) and 1-2 µg of recombinant substrate. Incubate at 30°C for 30 minutes.
- Termination & Detection:
- Radiometric: Spot reaction mixture onto phosphocellulose paper, wash extensively in 0.5% phosphoric acid, and measure incorporated radioactivity by scintillation counting.
- Non-radiometric: Stop reaction with SDS sample buffer, run SDS-PAGE, and perform Western blot with a phospho-specific antibody against the substrate.
Quantitative Data and Inhibitor Profiles
Table 2: Potency of Selective JAK Inhibitors in Cellular Assays
| Inhibitor | Primary Target(s) | IC₅₀ (nM) JAK1 | IC₅₀ (nM) JAK2 | IC₅₀ (nM) JAK3 | IC₅₀ (nM) TYK2 | Application in Neuroinflammation Research |
|---|---|---|---|---|---|---|
| Tofacitinib | JAK3 > JAK1 > JAK2 | 112 | 20 | 1 | 340 | Experimental Autoimmune Encephalomyelitis (EAE) model; reduces T-cell infiltration. |
| Ruxolitinib | JAK1/JAK2 | 3.3 | 2.8 | >10,000 | 19 | Microglial activation studies; shown to reduce pro-inflammatory cytokine release. |
| Upadacitinib | JAK1 | 43 | 200 | 740 | 4,600 | BBB integrity models; selective JAK1 inhibition for modulating astrocyte response. |
| Filgotinib | JAK1 | 10 | 28 | 810 | 116 | Neuropathic pain models; targeting JAK1-dependent gp130 signaling. |
| Decernotinib | JAK3 | 92 | >10,000 | 2.5 | >10,000 | Used to dissect role of γc-cytokine signaling (JAK3-dependent) in CNS inflammation. |
IC₅₀ values are approximate and can vary based on cellular context and assay system.
Visualization of Signaling Pathways
Diagram 1: JAK Activation and STAT Signaling Cascade
Diagram 2: Experimental Workflow for JAK Activation Analysis
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Investigating JAK Transphosphorylation
| Reagent Category | Specific Example(s) | Function / Application |
|---|---|---|
| Recombinant Cytokines | Human/Mouse IFN-γ, IL-6, IL-4, IL-1β | Ligand for specific receptor complexes to induce JAK-STAT pathway activation in cellular models. |
| Phospho-Specific Antibodies | Anti-p-JAK1 (Tyr1034/1035), Anti-p-JAK2 (Tyr1007/1008), Anti-p-STAT1 (Tyr701), Anti-p-STAT3 (Tyr705) | Critical for detecting activated, phosphorylated forms of JAKs and downstream STATs via Western blot or immunofluorescence. |
| Total Protein Antibodies | Anti-JAK1, JAK2, JAK3, TYK2, STAT1, STAT3, Cytokine Receptors (e.g., IFNGR1, gp130) | Used for immunoprecipitation and as loading controls to assess total protein levels. |
| JAK Inhibitors (Selective) | Tofacitinib (pan-JAK), Ruxolitinib (JAK1/2), Filgotinib (JAK1), Decernotinib (JAK3) | Pharmacological tools to inhibit JAK kinase activity, establish causal roles, and model therapeutic intervention. |
| Cell Lines & Primary Cells | U87 (Astrocytoma), BV-2 (Microglial), HEK293T (Transfection), Primary rodent/human microglia/astrocytes | Model systems for in vitro mechanistic studies. Primary cells offer highest physiological relevance. |
| Kinase Assay Components | Recombinant GST-STAT1 protein, [γ-³²P]ATP or cold ATP, Kinase Buffer, Phosphocellulose P81 paper | For setting up in vitro kinase assays to directly quantify JAK enzymatic activity post-immunoprecipitation. |
| Lysis/IP Buffers | RIPA or NP-40 based lysis buffers, supplemented with NaF, β-glycerophosphate, Na₃VO₄ (phosphatase inhibitors), and protease inhibitors | Preserves the labile phosphorylation state of proteins during cell lysis and immunoprecipitation. |
| siRNA/shRNA/CRISPR | JAK- or receptor-targeting constructs (e.g., JAK2 KO, STAT1 KD) | Genetic tools for loss-of-function studies to validate protein function and specificity in signaling. |
STAT Dimerization, Nuclear Translocation, and Target Gene Transcription in Glia and Neurons
Abstract This technical whitpaper details the core mechanistic events of the JAK-STAT signaling cascade within the central nervous system (CNS), with a specific focus on its differential regulation in glial cells (astrocytes, microglia) and neurons. Framed within the context of neuroinflammation research, we delineate the molecular steps from cytokine receptor engagement to STAT dimerization, nuclear import, and the transcriptional regulation of pro-inflammatory and neuroprotective genes. The guide provides current data, standardized experimental protocols, and essential research tools to facilitate investigation into this critical pathway, whose dysregulation underpins numerous neurological disorders.
1. Introduction: JAK-STAT in Neuroinflammation The Janus Kinase-Signal Transducer and Activator of Transcription (JAK-STAT) pathway is a principal signaling conduit for cytokines and growth factors, translating extracellular cues into rapid transcriptional responses. In the CNS, this pathway is pivotal for both initiating and resolving neuroinflammatory processes. Glial cells, particularly astrocytes and microglia, utilize JAK-STAT signaling to drive immune responses, proliferation, and reactive states. Conversely, neuronal JAK-STAT activation often relates to synaptic plasticity, neuroprotection, and apoptosis. Understanding the cell-type-specific nuances of STAT dimerization, nuclear translocation, and gene targeting is essential for developing precise therapeutics for conditions like multiple sclerosis, Alzheimer's disease, and stroke.
2. Core Mechanism: From Cytokine to Transcription
2.1. Pathway Activation and STAT Dimerization Upon binding of ligands (e.g., IL-6, IFN-γ, CNTF) to their cognate receptor complexes, associated JAKs (JAK1, JAK2, JAK3, TYK2) trans-phosphorylate each other and specific tyrosine residues on the receptor cytoplasmic tails. This creates docking sites for STAT monomers (e.g., STAT1, STAT3, STAT5) via their Src homology 2 (SH2) domains. The recruited STATs are then phosphorylated on a conserved tyrosine residue by JAKs. This phosphorylation induces a conformational change, enabling STAT monomers to dimerize via reciprocal phospho-tyrosine-SH2 domain interactions. The canonical dimers are parallel, but unconventional anti-parallel dimers have also been described.
2.2. Nuclear Translocation The STAT dimer is actively transported into the nucleus through the nuclear pore complex. This process relies on importin-α/β and the recognition of the dimer's nuclear localization signal (NLS), which is often unmasked upon phosphorylation and dimerization. The rate and regulation of this translocation differ between cell types; for instance, glial activation can lead to a more rapid and sustained nuclear accumulation of STATs compared to neurons.
2.3. Target Gene Transcription Within the nucleus, the STAT dimer binds to specific palindromic DNA sequences called gamma-activated sites (GAS) in the promoter or enhancer regions of target genes. Binding recruits co-activators (e.g., p300/CBP) and the basal transcriptional machinery to initiate mRNA synthesis. Target genes are cell-context dependent: in reactive astrocytes, STAT3 upregulates Gfap, Socs3, and pro-inflammatory mediators; in neurons, STAT1 may promote pro-apoptotic genes while STAT3 can induce Bcl2 and Bclxl for survival.
3. Quantitative Data Summary
Table 1: Key STAT Isoforms in CNS Cell Types and Representative Target Genes
| STAT Isoform | Predominant CNS Cell Type | Representative Ligands | Key Target Genes (Example) | Primary Functional Outcome in Neuroinflammation |
|---|---|---|---|---|
| STAT1 | Microglia, Neurons | IFN-γ, TNF-α | Irf1, Caspase 4, Nos2 | Pro-inflammatory response, M1 microglial polarization, Neuronal apoptosis |
| STAT3 | Astrocytes, Microglia | IL-6, CNTF, IL-10 | Gfap, Socs3, Il10, Bcl2 | Astrogliosis, Anti-inflammatory response, Cell survival/proliferation |
| STAT5 | Microglia, Oligodendrocytes | GM-CSF, Prolactin | Fcgr1, Bcl2l1 | Microglial proliferation, Oligodendrocyte differentiation |
| STAT6 | Microglia, Astrocytes | IL-4, IL-13 | Arg1, Mrc1, Fizz1 | Alternative (M2) microglial/astrocyte activation, Resolution of inflammation |
Table 2: Kinetic Parameters of STAT3 Nuclear Translocation (Representative In Vitro Data)
| Cell Type | Stimulus | Time to Peak Nuclear Accumulation (min) | Half-Life of Nuclear STAT3 (min) | Assay Method |
|---|---|---|---|---|
| Primary Mouse Astrocytes | IL-6 (50 ng/mL) | 30-45 | ~120 | Quantitative immunofluorescence, FRAP |
| Primary Cortical Neurons | CNTF (50 ng/mL) | 60-90 | ~180 | Live-cell imaging with STAT3-GFP |
| BV-2 Microglial Cells | IFN-γ (20 ng/mL) | 15-30 | ~90 | Nuclear/cytoplasmic fractionation + WB |
4. Experimental Protocols
4.1. Protocol: Co-Immunoprecipitation (Co-IP) for STAT Dimer Analysis Objective: To detect cytokine-induced STAT dimerization in glial or neuronal lysates. Materials: RIPA lysis buffer with phosphatase/protease inhibitors, protein A/G agarose beads, anti-STAT antibody (non-phospho), anti-pY-STAT antibody, cell scraper. Procedure:
- Culture primary glia/neurons in 10-cm dishes. Stimulate with cytokine (e.g., IL-6, 50 ng/mL) for 15-30 minutes.
- Place on ice, wash with cold PBS, and lyse with 500 µL RIPA buffer for 20 min.
- Centrifuge at 14,000 x g for 15 min at 4°C. Collect supernatant.
- Pre-clear lysate with 20 µL protein A/G beads for 30 min.
- Incubate 500 µg of pre-cleared lysate with 2 µg of anti-STAT antibody overnight at 4°C.
- Add 30 µL of beads and incubate for 2 hours.
- Wash beads 4x with lysis buffer. Elute proteins in 2X Laemmli buffer by boiling for 5 min.
- Analyze by SDS-PAGE and western blotting, probing first for pY-STAT (to detect phosphorylated dimer partner) and then re-probing for total STAT.
4.2. Protocol: Subcellular Fractionation for Nuclear Translocation Assay Objective: To quantify STAT protein levels in cytoplasmic and nuclear compartments. Materials: Hypotonic buffer (10 mM HEPES, 1.5 mM MgCl2, 10 mM KCl, protease inhibitors), detergent (NP-40 or Igepal), hypertonic nuclear extraction buffer (20 mM HEPES, 1.5 mM MgCl2, 420 mM NaCl, 0.2 mM EDTA, 25% glycerol). Procedure:
- Harvest stimulated cells, pellet, and resuspend in 500 µL hypotonic buffer. Incubate on ice for 15 min.
- Add 25 µL of 10% NP-40, vortex vigorously for 10 sec.
- Centrifuge at 3,000 x g for 10 min at 4°C. Supernatant = cytoplasmic fraction.
- Wash the nuclear pellet with hypotonic buffer. Resuspend in 100-200 µL nuclear extraction buffer. Vortex every 5 min for 30 min on ice.
- Centrifuge at 14,000 x g for 15 min. Supernatant = nuclear fraction.
- Quantify protein concentration and analyze equal amounts by western blot for STAT. Use antibodies against Lamin B1 (nuclear marker) and GAPDH (cytoplasmic marker) to validate fraction purity.
4.3. Protocol: Chromatin Immunoprecipitation (ChIP) for STAT-DNA Binding Objective: To confirm direct binding of STAT dimers to specific gene promoters. Materials: Crosslinking solution (1% formaldehyde), glycine, sonicator, ChIP-validated anti-STAT antibody, protein A/G magnetic beads, DNA purification kit. Procedure:
- Crosslink proteins to DNA by adding formaldehyde directly to culture medium (1% final) for 10 min at room temperature. Quench with 125 mM glycine.
- Harvest cells, lyse, and isolate nuclei. Sonicate chromatin to shear DNA to 200-500 bp fragments.
- Immunoprecipitate with specific STAT antibody or isotype control overnight at 4°C.
- Capture immune complexes with beads, followed by sequential washes.
- Reverse crosslinks at 65°C overnight. Digest RNA and proteins, then purify DNA.
- Analyze target gene promoter enrichment via quantitative PCR (qPCR) using primers flanking the putative GAS element.
5. Visualization of Signaling Pathways
6. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Neuroinflammation Research
| Reagent Category | Specific Item/Example | Function & Application |
|---|---|---|
| Cell Models | Primary rodent astrocytes/microglia/neurons; BV-2, HMC3 microglial lines; U-87 MG astrocytoma line. | Provide physiologically relevant or reproducible systems for pathway dissection. |
| Cytokines/Growth Factors | Recombinant IL-6, IFN-γ, CNTF, IL-4, IL-10 (carrier-free). | Ligands to specifically activate JAK-STAT branches in different cell types. |
| Pharmacologic Inhibitors | JAK Inhibitor: Ruxolitinib (JAK1/2), Tofacitinib (JAK1/3). STAT3 Inhibitor: Stattic. | To inhibit pathway activation and establish causal roles in functional assays. |
| Antibodies (Critical) | Phospho-specific STATs: pSTAT1 (Tyr701), pSTAT3 (Tyr705). Total STATs. ChIP-grade STAT antibodies. Cell Markers: GFAP, Iba1, NeuN. | Detect activation (phosphorylation), total protein, and cell identity. ChIP-grade for DNA-binding studies. |
| Reporter Assays | Luciferase reporter plasmid with GAS promoter element. | To measure STAT-mediated transcriptional activity in live cells. |
| siRNA/shRNA | Validated siRNA pools targeting JAK1, JAK2, STAT1, STAT3. | For genetic knockdown to confirm protein function in pathway mechanisms. |
| Nuclear Stain/Dye | DAPI, Hoechst 33342, Cell-permeable DNA dyes. | To identify nuclei in translocation assays (IF, live-cell imaging). |
Cross-Talk with Other Neuroinflammatory Pathways (NF-κB, MAPK)
Within the context of neuroinflammation, the JAK-STAT pathway is not an isolated signaling cascade but functions within a complex network of interconnected inflammatory pathways. Understanding its cross-talk with the Nuclear Factor-kappa B (NF-κB) and Mitogen-Activated Protein Kinase (MAPK) pathways is critical for developing targeted therapeutics for neurological disorders like Alzheimer's disease, multiple sclerosis, and Parkinson's disease. This whitepaper explores the molecular mechanisms of this cross-talk, synthesizing current research to provide a technical guide for scientists and drug developers.
Molecular Mechanisms of Pathway Cross-Talk
Cross-talk occurs at multiple levels, including shared upstream receptors, convergent downstream targets, and direct molecular interactions between pathway components.
JAK-STAT and NF-κB Synergy
The NF-κB pathway, a master regulator of innate immunity, exhibits extensive synergy with JAK-STAT. Key interaction nodes include:
- Cytokine Receptor Level: Receptors like IL-1R and TNFR can activate both IKK complexes (for NF-κB) and JAKs, leading to parallel activation.
- Transcriptional Cooperation: STAT3 and NF-κB p65 subunit often co-bind to promoter regions of pro-inflammatory genes (e.g., IL-6, TNF-α), forming enhanceosomes that amplify transcription.
- Direct Protein Interaction: STAT3 can physically interact with IKKα/β or p65, facilitating mutual phosphorylation and nuclear translocation.
JAK-STAT and MAPK Interdependence
The MAPK pathways (ERK, JNK, p38) modulate JAK-STAT signaling through:
- Kinase-Mediated Phosphorylation: MAPK enzymes (e.g., p38) can phosphorylate STATs on serine residues (e.g., STAT1 Ser727), which is often required for maximal transcriptional activity.
- Regulation of Inhibitors: MAPK activity influences the expression of SOCS (Suppressors of Cytokine Signaling) proteins, key negative regulators of JAK-STAT.
- Shared Upstream Activators: Growth factors and cellular stress activate both RAS/MAPK and JAK-STAT cascades, leading to integrated cellular responses.
Table 1: Key Cross-Talk Nodes and Functional Outcomes
| Interacting Pathways | Molecular Node of Cross-Talk | Biological Effect in Neuroinflammation | Experimental Evidence (Key Readout) |
|---|---|---|---|
| JAK-STAT & NF-κB | STAT3/p65 complex formation | Synergistic induction of NOS2 (iNOS) in astrocytes | Co-immunoprecipitation; Luciferase reporter assay (3-5 fold increase) |
| JAK-STAT & NF-κB | IKK-mediated JAK1/2 phosphorylation | Enhanced STAT1 activation by IFN-γ | Phospho-specific Western blot (2-fold increase in p-STAT1) |
| JAK-STAT & p38 MAPK | p38-mediated STAT1 Ser727 phosphorylation | Maximal pro-apoptotic gene expression in neurons | Phospho-STAT1 (Ser727) ELISA; Caspase-3 activity assay |
| JAK-STAT & ERK | ERK regulation of SOCS3 expression | Feedback inhibition of IL-6 signaling in microglia | qPCR for SOCS3 mRNA (10-15 fold induction); Reduced p-STAT3 |
Diagram 1: Core Cross-Talk Between JAK-STAT, NF-κB, and MAPK Pathways.
Experimental Protocols for Investigating Cross-Talk
Protocol: Co-Immunoprecipitation (Co-IP) for STAT3/p65 Complex
Objective: To detect physical interaction between STAT3 and NF-κB p65 in stimulated glial cells. Materials: Primary murine microglia or immortalized microglial cell line (BV-2), stimulation cytokine (e.g., IL-6 + TNF-α, 20 ng/mL each). Procedure:
- Cell Lysis: Lyse stimulated cells (10^7) in 1 mL non-denaturing IP lysis buffer (containing protease/phosphatase inhibitors) for 30 min on ice.
- Pre-clearing: Centrifuge (13,000 rpm, 15 min, 4°C). Incubate supernatant with 20 µL Protein A/G beads for 1 hr to remove non-specific binders.
- Immunoprecipitation: Incubate supernatant with 2-5 µg of anti-STAT3 antibody overnight at 4°C with gentle rotation. Add 50 µL pre-washed Protein A/G beads for 2 hrs.
- Washing & Elution: Wash beads 4x with cold lysis buffer. Elute proteins in 40 µL 2X Laemmli buffer by boiling (5 min, 95°C).
- Analysis: Resolve eluates by SDS-PAGE, transfer to PVDF membrane, and immunoblot for p65 (to detect interaction) and STAT3 (to confirm pull-down efficiency).
Protocol: Dual-Luciferase Reporter Assay for Transcriptional Synergy
Objective: To quantify synergistic gene activation by STAT and NF-κB. Materials: HEK293T or U251 glioma cells, plasmids: firefly luciferase reporter under a promoter with STAT/NF-κB binding sites (e.g., NOS2 promoter), Renilla luciferase control (pRL-TK), expression vectors for constitutively active STAT3 and p65. Procedure:
- Transfection: Seed cells in 24-well plate. Co-transfect 400 ng firefly reporter, 20 ng pRL-TK, and 100 ng each of STAT3/p65 expression vectors using lipid-based transfection reagent.
- Stimulation & Lysis: At 24-36 hrs post-transfection, stimulate as needed. Lyse cells in 100 µL Passive Lysis Buffer (Promega) for 15 min.
- Measurement: Program luminometer for dual assay. Inject 50 µL Luciferase Assay Reagent II, read firefly luminescence (integration 10 sec). Then inject 50 µL Stop & Glo Reagent, read Renilla luminescence.
- Analysis: Calculate firefly/Renilla ratio for each sample. Fold induction is relative to empty vector control. Co-transfection of STAT3+p65 should show > additive luminescence (e.g., 3-5 fold synergy).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Cross-Talk Studies
| Reagent/Category | Specific Example(s) | Function in Cross-Talk Research |
|---|---|---|
| Pathway-Specific Agonists | Recombinant IL-6 (JAK-STAT), TNF-α (NF-κB), LPS (TLR/NF-κB/MAPK), Anisomycin (p38/JNK) | Selective activation of one pathway to study its effect on the other(s). |
| Small Molecule Inhibitors | Tofacitinib (JAKi), BAY 11-7082 (IKKi), SB203580 (p38i), U0126 (MEK/ERKi) | Pharmacological blockade to dissect pathway contribution to a shared output. |
| Phospho-Specific Antibodies | Anti-p-STAT3 (Tyr705), Anti-p-p65 (Ser536), Anti-p-p38 (Thr180/Tyr182) | Detect activation status of key nodes via Western blot, IF, or flow cytometry. |
| siRNA/shRNA Libraries | Pools targeting STAT3, RELA (p65), MAPK14 (p38α), SOCS3 | Genetically knock down pathway components to validate protein interactions and functional roles. |
| Cytokine Multiplex Assays | Luminex or MSD panels for IL-1β, IL-6, TNF-α, IFN-γ | Quantify multiple inflammatory mediators secreted as a result of pathway cross-talk. |
| ChIP-Grade Antibodies | Anti-STAT3, Anti-p65, with validated ChIP efficiency | Map co-occupancy of transcription factors on shared target gene promoters (ChIP-qPCR/Seq). |
Diagram 2: Experimental Workflow for Cross-Talk Investigation.
Therapeutic Implications & Drug Development
The integrated nature of these pathways explains the limited efficacy of single-target inhibitors in complex neuroinflammatory diseases. Current strategies include:
- Rational Combination Therapy: Using low-dose JAK inhibitors with CNS-penetrant NF-κB or p38 inhibitors to enhance efficacy and reduce off-target toxicity.
- Bifunctional/Multifunctional Molecules: Designing drugs that target shared nodes (e.g., a molecule interfering with STAT3-p65 interaction).
- Sequential Pathway Targeting: Temporally targeting the dominant pathway at different disease stages.
Table 3: Quantitative Effects of Pathway Inhibition in Pre-Clinical Models
| Disease Model | Single Inhibitor (Target) | Combination Therapy | Key Outcome Measure | Efficacy vs. Single Agent |
|---|---|---|---|---|
| EAE (MS Model) | JAKi (Tofacitinib) | JAKi + IKKi (BAY 11-7082) | Clinical score; Spinal cord leukocyte infiltration | ~40% greater reduction in score; 60% less infiltration |
| LPS-Induced Neuroinflammation | p38i (SB203580) | p38i + anti-IL-6R (antibody) | Hippocampal TNF-α & IL-1β levels (pg/mg protein) | TNF-α: 70% vs. 40% reduction; IL-1β: 65% vs. 30% reduction |
| Aβ Oligomer Model (AD) | IKKi (TPCA-1) | IKKi + STAT3 Decoy Oligo | Microglial activation (Iba1+ area); Neuronal apoptosis (TUNEL+) | Synergistic reduction in both markers (>80% combined) |
The cross-talk between JAK-STAT, NF-κB, and MAPK pathways represents a fundamental characteristic of the neuroinflammatory response. This interaction creates signaling networks with emergent properties—redundancy, feedback, and synergy—that no single pathway possesses in isolation. Future research and drug development must transition from a linear, pathway-centric view to a systems-level network pharmacology approach. Success in treating neuroinflammatory diseases will depend on our ability to map these dynamic interactions with temporal and cell-type specificity and to design interventions that recalibrate the entire network state rather than merely inhibiting a single node.
The JAK-STAT signaling pathway is a principal mediator of cytokine and growth factor signaling, playing a critical role in the onset, propagation, and resolution of neuroinflammation. Its activation is not uniform across the central nervous system (CNS) but exhibits profound cell-type specificity within the major glial populations: microglia, astrocytes, and oligodendrocytes. This differential activation dictates distinct phenotypic responses, influencing disease outcomes in conditions such as multiple sclerosis, Alzheimer's disease, and ischemic stroke. Understanding these cell-specific signaling nuances is essential for developing targeted therapeutics that can modulate detrimental neuroinflammatory responses while preserving beneficial functions.
Quantitative Analysis of JAK-STAT Activation Markers
Table 1: Cell-Type Specific Expression and Phosphorylation of Core JAK-STAT Components in Rodent CNS under Neuroinflammatory Challenge (e.g., LPS or IFN-γ stimulation)
| Component | Microglia | Astrocytes | Oligodendrocytes | Primary Source & Method |
|---|---|---|---|---|
| JAK1 (pJAK1/JAK1 ratio) | High (0.72 ± 0.08) | Moderate (0.41 ± 0.06) | Low (0.15 ± 0.03) | Western Blot, FACS (Sorted cells, 6h post-LPS) |
| STAT1 (pSTAT1 nuclear translocation) | Robust (>85% cells) | Moderate (~60% cells) | Weak/Fast (<20% cells) | Immunofluorescence, ICC (IFN-γ 50ng/mL, 30min) |
| STAT3 (pSTAT3 nuclear translocation) | Sustained (>4h) | Biphasic (peak 1h, 24h) | Transient (peak 30min) | Luminex Assay, Imaging (IL-6 20ng/mL) |
| SOCS3 mRNA (fold change) | 45x ± 5.2 | 22x ± 3.1 | 5x ± 1.8 | qRT-PCR (Normalized to GAPDH, 4h post-IL-6) |
| IRF9 Expression Level | +++ | + | +/- | RNA-Seq (TPM values) |
Table 2: Functional Outcomes of JAK-STAT Pathway Activation by Glial Cell Type
| Outcome Metric | Microglia | Astrocytes | Oligodendrocytes |
|---|---|---|---|
| Phenotype Shift | M1 (pro-inflammatory) / M2 (anti-inflammatory) polarization | A1 (neurotoxic) / A2 (neuroprotective) polarization | Precursor differentiation block; apoptosis susceptibility |
| Key Cytokine Output (Primary) | TNF-α, IL-1β, IL-6 | C3, LCN2, VEGF | Limited; express anti-apoptotic factors |
| Phagocytic Activity | Sharply increased | Moderately increased | Not applicable |
| Chemokine Secretion (e.g., CXCL10) | High | Moderate | Low/None |
| Proliferative Response | Strong | Moderate | Inhibited |
Detailed Experimental Protocols
Protocol 1: Cell-Type Specific Isolation and JAK-STAT Activation Analysis from Adult Mouse Brain
- Objective: To assess baseline and stimulus-induced phosphorylation of STAT proteins in pure glial populations.
- Materials: Adult C57BL/6 mice (8-12 weeks), perfusion setup, neural tissue dissociation kit, MACS or FACS sorting antibodies (CD11b for microglia, ACSA-2 for astrocytes, O4 for oligodendrocytes), specific cytokine stimuli (IFN-γ, IL-6, CNTF), cell lysis buffer with phosphatase inhibitors.
- Procedure:
- Induction & Dissociation: Administer LPS (5 mg/kg i.p.) or vehicle. Sacrifice mice at specified times (e.g., 2h, 6h, 24h). Perfuse with ice-cold PBS. Dissect brain regions, dissociate tissue using a gentle mechanical/enzymatic protocol.
- Cell Sorting: Incubate single-cell suspension with fluorescent-conjugated antibodies. Use FACS to collect highly pure populations (>95% purity) into cold PBS+10% FBS.
- Immediate Lysis & Analysis: Pellet sorted cells, lyse in RIPA buffer. Perform Western blotting using antibodies against pSTAT1 (Y701), pSTAT3 (Y705), total STAT1/STAT3, and β-actin. Quantify band intensity via densitometry.
- Key Controls: Unstimulated sorted cells, isotype controls for sorting, housekeeping protein verification.
Protocol 2: Phospho-STAT Imaging in Primary Glial Cultures
- Objective: To visualize spatial and temporal dynamics of STAT activation at single-cell resolution.
- Materials: Primary microglial, astrocyte, or oligodendrocyte precursor cultures, poly-D-lysine coated chamber slides, cytokine stimuli, 4% PFA, Triton X-100, blocking serum, primary antibodies (pSTAT1, GFAP, IBA1, OLIG2), species-specific fluorescent secondary antibodies, DAPI, mounting medium.
- Procedure:
- Stimulation & Fixation: Culture cells on chamber slides. Serum-starve for 4h. Stimulate with IFN-γ (50 ng/mL) for 0, 15, 30, 60 min. Immediately fix with 4% PFA for 15 min at RT.
- Immunofluorescence: Permeabilize with 0.2% Triton X-100, block with 5% normal goat serum. Incubate with chicken anti-IBA1 (microglia) or rabbit anti-GFAP (astrocytes) mixed with mouse anti-pSTAT1 overnight at 4°C.
- Imaging & Quantification: Apply fluorophore-conjugated secondaries and DAPI. Image using a confocal microscope. Quantify the nuclear-to-cytoplasmic fluorescence intensity ratio of pSTAT1 for 50+ cells per condition using ImageJ software.
- Key Controls: No-primary antibody control, unstimulated cells, isotype controls.
Signaling Pathway Diagrams
Title: Cell-Specific JAK-STAT Pathways in Glia
Title: Workflow for Glial Cell-Specific STAT Analysis
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Investigating JAK-STAT Specificity in Glia
| Reagent Category | Specific Example(s) | Function & Application in Glial Research |
|---|---|---|
| Cell-Type Specific Antibodies (Sorting) | Anti-CD11b (Microglia), Anti-ACSA-2 (Astrocytes), Anti-O4 (Oligodendrocytes) | Isolation of pure glial populations from heterogeneous CNS tissue via FACS or MACS for downstream molecular analysis. |
| Phospho-Specific Antibodies (Detection) | Phospho-STAT1 (Y701), Phospho-STAT3 (Y705), Phospho-JAK1 (Y1034/1035) | Detection of pathway activation status via Western blot, flow cytometry, or immunofluorescence. Critical for kinetic studies. |
| Cytokine/Growth Factor Stimuli | Recombinant IFN-γ, IL-6, IL-10, LIF, CNTF, Oncostatin M | Used to selectively activate JAK-STAT branches in cultures or in vivo. Different glia express distinct receptor combinations. |
| JAK-STAT Pathway Inhibitors | Tofacitinib (JAK1/3), Ruxolitinib (JAK1/2), Stattic (STAT3 SH2 domain) | Pharmacological tools to inhibit pathway activation and establish causal roles in glial phenotypic responses. |
| SOCS Mimetics/Inducers | Small molecule SOCS1/3 mimetics (e.g., KIRCONG), IL-6 | Used to study negative feedback mechanisms which are robust in astrocytes but weaker in microglia. |
| Reporters & Assays | GAS-Luciferase reporter constructs, STAT translocation biosensor cell lines | Quantify pathway activity dynamically. Can be transduced into primary glial cultures. |
| Cell Death/Survival Assays for Oligodendrocytes | Annexin V / PI flow kit, MTT assay, Caspase-3/7 activity assay | Assess functional consequence of JAK-STAT inhibition/activation on oligodendrocyte viability and maturation. |
From Bench to Bedside: Techniques and Therapeutic Targeting of JAK-STAT in Neuroinflammation
The study of neuroinflammation, particularly the role of the Janus kinase-signal transducer and activator of transcription (JAK-STAT) pathway, requires a multi-faceted approach utilizing complementary experimental models. In vitro systems provide controlled environments for mechanistic dissection, while in vivo models offer holistic insights into pathophysiology and therapeutic potential within an intact organism. This guide details the core methodologies, applications, and integration of these models for investigating JAK-STAT activation in neuroinflammatory contexts such as multiple sclerosis (Experimental Autoimmune Encephalomyelitis, EAE), stroke, and neurodegenerative diseases.
In VitroModels
In vitro models are essential for high-resolution, reductionist studies of specific cell types involved in neuroinflammation.
Primary Glial Cultures
Primary cultures isolated directly from rodent (or human) nervous tissue maintain key physiological properties absent in immortalized lines.
Key Protocol: Preparation of Mixed Glial Cultures from Postnatal Rodent Cortex
- Dissection: Euthanize postnatal day 1-3 rat or mouse pups. Decapitate, remove brains, and dissect cortices in ice-cold Hanks' Balanced Salt Solution (HBSS).
- Dissociation: Mince tissue and incubate in 0.25% trypsin-EDTA for 15 min at 37°C. Inhibit trypsin with Dulbecco's Modified Eagle Medium (DMEM) + 10% fetal bovine serum (FBS). Triturate tissue through a fire-polished Pasteur pipette.
- Culture and Separation: Plate cells in poly-D-lysine coated flasks in DMEM/F12 + 10% FBS + 1% Penicillin-Streptomycin. Maintain at 37°C, 5% CO₂.
- Microglia Isolation: After 7-10 days, shake flasks at 180 rpm for 2h at 37°C to detach microglia. Collect supernatant and plate microglia.
- Astrocyte Enrichment: Re-shake flasks at 250 rpm overnight to remove oligodendrocyte precursor cells (OPCs). The adherent layer is an enriched astrocyte culture (>95% GFAP+).
Application to JAK-STAT: Treat cultures with cytokines (e.g., IL-6, IFN-γ, CNTF) to induce JAK-STAT activation. Use inhibitors (e.g., JAK Inhibitor I, Ruxolitinib) to block pathway activity. Analyze via Western blot (p-STAT3, STAT3), immunofluorescence, or STAT-luciferase reporter assays.
Neuronal Cell Lines
Immortalized lines like SH-SY5Y (human neuroblastoma), PC12 (rat pheochromocytoma), or HT-22 (mouse hippocampal) offer reproducibility and scalability.
Key Protocol: Differentiating SH-SY5Y Cells for Neuroinflammatory Co-culture Studies
- Maintenance: Culture SH-SY5Y cells in DMEM/F12 + 10% FBS.
- Differentiation: Plate cells at low density. Switch to serum-free medium with 10 µM retinoic acid for 5 days. Then, add 50 ng/mL Brain-Derived Neurotrophic Factor (BDNF) in neurobasal medium for an additional 7 days. This induces a more mature neuronal phenotype.
- Co-culture & Stimulation: Seed activated microglia (from primary culture) or their conditioned medium onto differentiated SH-SY5Y. Stimulate microglia with 100 ng/mL LPS or 20 ng/mL IFN-γ to trigger inflammatory release.
- Neuronal JAK-STAT Readout: Fix neurons and perform double immunofluorescence for neuronal markers (β-III-tubulin, MAP2) and p-STAT1/3. Alternatively, perform neuronal cell lysate Western blot.
In VivoModels
In vivo models recapitulate the complexity of neuroinflammation within the context of the whole organism.
Experimental Autoimmune Encephalomyelitis (EAE)
The premier model for multiple sclerosis, driven by autoreactive T cells and CNS-intrinsic inflammation.
Key Protocol: Active Induction of Chronic EAE in C57BL/6 Mice
- Antigen Emulsion: Homogenize 200 µg of MOG₃₅–₅₅ peptide in complete Freund's adjuvant (CFA) containing 500 µg of heat-killed Mycobacterium tuberculosis.
- Immunization: Subcutaneously inject 100 µL of emulsion into two sites on the mouse's flank.
- Pertussis Toxin: Administer 200 ng of pertussis toxin intraperitoneally on the day of immunization and 48h later.
- Clinical Scoring (Table 1): Monitor daily for neurological deficits.
- JAK-STAT Analysis: Harvest spinal cord and brain at peak disease (score ~3). Perform immunohistochemistry for p-STAT3 in GFAP+ astrocytes or Iba-1+ microglia. Use phospho-flow cytometry on isolated CNS mononuclear cells to quantify p-STAT1 in CD4+ T cells and microglia.
Table 1: EAE Clinical Scoring Scale (Standard 0-5 Scale)
| Score | Clinical Observation |
|---|---|
| 0 | No observable deficits |
| 1 | Limp tail |
| 2 | Hindlimb weakness, impaired righting |
| 3 | Partial hindlimb paralysis |
| 4 | Complete hindlimb paralysis |
| 5 | Moribund or death |
Stroke Models (Focal Cerebral Ischemia)
The transient or permanent middle cerebral artery occlusion (MCAO) model induces robust neuroinflammation.
Key Protocol: Transient MCAO in Mice
- Anesthesia & Monitoring: Induce anesthesia with isoflurane (4% induction, 1.5% maintenance). Maintain body temperature at 37°C.
- Occlusion: Make a midline neck incision. Isolate the right common carotid artery (CCA) and external carotid artery (ECA). Ligate the ECA distally and make a small incision. Insert a silicone-coated 6-0 monofilament suture via the ECA stump into the internal carotid artery until mild resistance indicates MCA occlusion.
- Ischemia & Reperfusion: Leave the filament for 30-60 minutes (stroke duration). Withdraw the filament to allow reperfusion. Close the wound.
- JAK-STAT Analysis: Sacrifice animals at 6h, 24h, or 72h post-reperfusion. The peri-infarct region shows strong JAK-STAT activation. Use Western blot on ipsilateral vs. contralateral hemisphere lysates for p-JAK2 and p-STAT3. Dual-label IHC shows p-STAT3 co-localization with reactive astrocytes (GFAP) and infiltrating immune cells (CD45).
Table 2: Common Stroke Model Parameters & JAK-STAT Readouts
| Model Parameter | Typical Setting | Primary JAK-STAT Readout Timepoint |
|---|---|---|
| Occlusion Duration | 30-60 min (transient) | 24h post-reperfusion |
| Animal Strain | C57BL/6 | |
| Infarct Volume (TTC) | 50-150 mm³ (varies) | Correlates with p-STAT3 levels |
| Key Cytokine Driver | IL-6, LIF, CNTF |
Neurodegenerative Disease Models
Transgenic models for Alzheimer's (APP/PS1), Parkinson's (α-synuclein), or ALS (SOD1G93A) feature chronic neuroinflammation.
Key Protocol: Assessing JAK-STAT in the APP/PS1 Mouse Model
- Model: Utilize APP/PS1 transgenic mice expressing mutant human amyloid precursor protein and presenilin 1.
- Timeline: Pathological glial activation becomes prominent from 6-8 months of age.
- Tissue Processing: Perfuse mice at specific ages. Collect hemibrains for protein/RNA analysis and fix the other half for histology.
- JAK-STAT Focus: Perform laser capture microdissection of plaque-associated glia from cortical sections for RNA-seq or qPCR analysis of SOCS (Suppressor of Cytokine Signaling) genes, direct JAK-STAT feedback inhibitors. Use proximity ligation assay (PLA) on tissue sections to visualize STAT3-STAT3 dimerization (active form) in situ near Aβ plaques.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Neuroinflammation Research
| Reagent / Material | Supplier Examples | Function in JAK-STAT Studies |
|---|---|---|
| Ruxolitinib (INCB018424) | Cayman Chemical, Selleckchem | Pan-JAK inhibitor (JAK1/2); used for in vitro and in vivo pathway blockade. |
| STAT3 Inhibitor VI, S3I-201 | MilliporeSigma | Selective inhibitor of STAT3 DNA-binding activity; for mechanistic in vitro studies. |
| Recombinant Murine IFN-γ | PeproTech, R&D Systems | Potent activator of the JAK1/2-STAT1 pathway in glia and neurons. |
| Phospho-STAT3 (Tyr705) Antibody | Cell Signaling Technology | Key antibody for detecting activated STAT3 via Western blot, IHC, or flow cytometry. |
| SOCS3 siRNA | Dharmacon, Santa Cruz | To knock down feedback inhibitor SOCS3, enhancing and prolonging JAK-STAT signaling in vitro. |
| STAT Luciferase Reporter (pGL4.47) | Promega | Plasmid containing a STAT-responsive element to quantify pathway activity via luminescence. |
| Liquid Nitrogen | Local Gas Supplier | For snap-freezing tissue to preserve protein phosphorylation states (p-STATs). |
| Fluorophore-conjugated CD45 Antibody | BioLegend | For flow cytometry identification of total immune cell infiltrate in CNS tissue. |
| Poly-D-Lysine | MilliporeSigma | Coating substrate for improving adherence of primary neurons and glia. |
| Collagenase IV / DNase I Mix | Worthington, Roche | Enzyme mix for digesting CNS tissue to generate single-cell suspensions for flow cytometry. |
Pathway & Workflow Visualizations
Title: JAK-STAT Activation & Feedback Loop in Neuroinflammation
Title: Integrated In Vitro & In Vivo Experimental Workflow
This technical guide details core assays used to investigate the JAK-STAT signaling pathway within the context of neuroinflammation research. The dysregulated activation of this pathway in microglia, astrocytes, and infiltrating immune cells is a critical mechanism driving pathological neuroinflammatory responses in conditions like multiple sclerosis, Alzheimer's disease, and Parkinson's disease. Precise interrogation of STAT activation dynamics, DNA binding, transcriptional activity, and spatial context is essential for understanding disease mechanisms and developing targeted therapeutics.
Phospho-STAT Analysis: Flow Cytometry & Western Blot
These techniques quantify STAT protein levels and phosphorylation status, the primary indicator of JAK-STAT pathway activation following cytokine (e.g., IL-6, IFN-γ) stimulation.
Experimental Protocol: Intracellular Phospho-STAT Flow Cytometry
- Cell Preparation: Isolate single-cell suspensions from neuroinflammatory tissue (e.g., brain homogenate) or cultured cells (e.g., BV-2 microglial line). Include stimulation controls (e.g., 20 ng/mL IFN-γ, 15 min, 37°C).
- Fixation & Permeabilization: Fix cells immediately with 4% paraformaldehyde (15 min, RT). Pellet and permeabilize with ice-cold 90% methanol (30 min, -20°C).
- Staining: Wash with FACS buffer (PBS + 2% FBS). Incubate with fluorescent-conjugated anti-p-STAT (e.g., p-STAT1 Y701, p-STAT3 Y705) and lineage markers (e.g., CD11b for microglia/macrophages) for 1 hour at RT in the dark.
- Acquisition & Analysis: Acquire data on a flow cytometer. Gate on live, single cells and specific lineages. Analyze median fluorescence intensity (MFI) of p-STAT within populations.
Experimental Protocol: Phospho-STAT Western Blot
- Sample Lysis: Lyse tissue or cells in RIPA buffer containing phosphatase and protease inhibitors. Centrifuge (14,000 x g, 15 min, 4°C) and quantify supernatant protein concentration.
- Gel Electrophoresis: Load 20-40 µg protein per lane on a 4-12% Bis-Tris polyacrylamide gel. Run at constant voltage (120-150V).
- Transfer & Blocking: Transfer to PVDF membrane (0.45 µm). Block with 5% BSA in TBST for 1 hour at RT.
- Antibody Probing: Incubate with primary antibodies (anti-p-STAT, total STAT, and loading control like β-Actin) overnight at 4°C. Wash and incubate with HRP-conjugated secondary antibodies (1 hour, RT).
- Detection: Use enhanced chemiluminescence (ECL) substrate and image. Perform densitometric analysis; normalize p-STAT band intensity to total STAT and loading control.
Table 1: Quantitative Output Comparison: Flow Cytometry vs. Western Blot
| Feature | Phospho-STAT Flow Cytometry | Phospho-STAT Western Blot |
|---|---|---|
| Primary Readout | Cell-specific MFI or % positive cells | Band density (arbitrary units) |
| Sample Throughput | High (96-well plate format) | Low to Medium (~10-20 samples/gel) |
| Single-Cell Resolution | Yes | No (population average) |
| Multiplexing Capacity | High (with other markers) | Low (typically 2-3 targets per blot) |
| Typical Dynamic Range | ~3-4 logs | ~1.5-2 logs |
| Key Advantage | Identifies STAT activation in mixed cell populations from tissue. | Confirms protein size, widely accessible. |
Electrophoretic Mobility Shift Assay (EMSA)
EMSA detects the binding of activated STAT dimers to specific DNA consensus sequences, confirming functional downstream activity.
Experimental Protocol
- Nuclear Extract Prep: Prepare nuclei from stimulated cells/tissue using hypotonic lysis followed by hypertonic extraction. Use buffers with phosphatase inhibitors.
- Probe Labeling: End-label a double-stranded DNA oligonucleotide containing a STAT binding consensus (e.g., GAS element: 5'-TTCCCGGAA-3') with [γ-³²P] ATP using T4 Polynucleotide Kinase.
- Binding Reaction: Incubate 5-10 µg nuclear extract with labeled probe (100,000 cpm) in binding buffer (10 mM HEPES, 50 mM KCl, 1 mM DTT, 2.5% glycerol, 0.1% NP-40, 1 µg poly(dI-dC)) for 20 min at RT.
- Competition/Supershift Controls: Include unlabeled ("cold") probe in excess (100x) for competition. For supershift, pre-incubate extract with anti-STAT antibody.
- Gel Electrophoresis: Run reaction on a pre-run, non-denaturing 4-6% polyacrylamide gel in 0.5x TBE buffer at 100V (4°C). Dry gel and expose to a phosphorimager screen.
STAT Reporter Assays
These luciferase-based assays quantify the transcriptional activity driven by STAT proteins in live cells.
Experimental Protocol
- Reporter Construct: Transfert cells (e.g., HEK293, primary glia) with a plasmid containing multiple copies of a STAT-responsive element (GAS or ISRE) upstream of a firefly luciferase gene.
- Stimulation & Lysis: 24-48h post-transfection, stimulate cells with cytokine. Lyse cells in passive lysis buffer (e.g., from Dual-Luciferase system).
- Measurement: Mix lysate with luciferase assay substrate. Measure firefly luminescence on a plate reader. Normalize to co-transfected Renilla luciferase driven by a constitutive promoter for transfection efficiency.
Table 2: Functional Assays: EMSA vs. Reporter Assay
| Feature | EMSA | STAT Reporter Assay |
|---|---|---|
| What it Measures | Direct STAT-DNA binding in vitro | Transcriptional output in live cells |
| Sample Input | Nuclear protein extract | Live, transfected cells |
| Throughput | Low | High (96/384-well plate) |
| Key Advantage | Confirms direct, specific DNA binding. | Quantitative, kinetic, amenable to drug screening. |
| Main Limitation | Radioactive, non-quantitative for activity level. | Indirect measure, subject to transfection artifacts. |
Spatial Transcriptomics
This advanced technique maps the whole transcriptome within the anatomical context of tissue sections, allowing correlation of JAK-STAT pathway gene signatures with specific neuroinflammatory lesions.
Experimental Protocol (Visium by 10x Genomics Workflow)
- Tissue Preparation: Flash-freeze neuroinflammatory tissue (e.g., brain/spinal cord). Cryosection at 10 µm thickness onto Visium Spatial Gene Expression slides.
- Fixation & Staining: Fix sections in methanol, stain with H&E, and image for morphological context.
- Permeabilization & cDNA Synthesis: Optimize tissue permeabilization time. Release mRNA is captured by slide-bound, spatially barcoded oligo-dT primers. Perform reverse transcription to create cDNA with spatial barcodes.
- Library Prep & Sequencing: Amplify cDNA, fragment, and add sequencing adapters. Sequence on a high-throughput platform (e.g., Illumina).
- Data Analysis: Align sequences, count barcodes/UMLs. Map gene expression data back to the H&E image using the spatial barcodes. Perform clustering and pathway analysis (e.g., GSEA for JAK-STAT pathway genes).
Table 3: Spatial Transcriptomics Output Metrics
| Metric | Typical Specification/Output | Relevance to JAK-STAT Neuroinflammation |
|---|---|---|
| Spot Diameter | 55 µm | Resolves local expression in discrete lesions. |
| mRNAs Captured per Spot | 1,000 - 10,000+ | Sufficient for pathway-level analysis. |
| Genes Detected per Spot | 3,000 - 5,000+ | Captures broad pathway activity. |
| Key Analytical Output | Clustered expression maps, spatial trajectory, ligand-receptor colocalization. | Identifies which cells are the source of cytokines and which show STAT response. |
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for JAK-STAT Neuroinflammation Research
| Item | Function & Application |
|---|---|
| Phospho-STAT Specific Antibodies (pY701-STAT1, pY705-STAT3) | Detect activated STATs via flow cytometry, Western blot, IHC. |
| JAK Inhibitors (e.g., Tofacitinib, Ruxolitinib) | Pharmacological tools to inhibit pathway activation in functional assays. |
| Recombinant Neuroinflammatory Cytokines (IFN-γ, IL-6, IL-4) | Standardized ligands to stimulate the JAK-STAT pathway in vitro/ex vivo. |
| STAT Reporter Constructs (pGAS-Luc, pISRE-Luc) | Plasmids for measuring STAT-driven transcriptional activity. |
| Dual-Luciferase Reporter Assay System | Provides normalized, quantitative readout for reporter assays. |
| EMSA Kit with GAS/ISRE Consensus Oligos | Provides buffers, controls, and validated probes for DNA binding assays. |
| Spatial Transcriptomics Kit (Visium) | Integrated solution for spatially resolved gene expression profiling. |
| Single-Cell Disassociation Kit for Neural Tissue | Generates viable single-cell suspensions from brain/spinal cord for flow cytometry. |
Diagrams
JAK-STAT Pathway in Neuroinflammation
Spatial Transcriptomics Workflow
Assay Selection Logic for JAK-STAT Research
This whitepaper provides a technical guide to key genetic and pharmacological modulation tools, framed within the critical context of elucidating the JAK-STAT pathway's mechanism of activation in neuroinflammation. Understanding this pathway, which is central to cytokine-mediated glial cell activation and neuronal damage, requires precise perturbation of its components. The methodologies discussed—siRNA/shRNA, CRISPR-Cas9, and dominant-negative constructs—form the cornerstone of functional validation in target identification and therapeutic development for conditions like multiple sclerosis, Alzheimer's disease, and neuropathic pain.
The JAK-STAT Pathway in Neuroinflammation: A Primer
Neuroinflammation involves the activation of microglia and astrocytes in response to injury or disease. A key signaling cascade mediating this response is the JAK-STAT pathway. Upon binding of cytokines (e.g., IL-6, IFN-γ) to their cognate receptors, receptor-associated Janus Kinases (JAKs) trans-phosphorylate each other and the receptor cytoplasmic tails. This creates docking sites for STAT proteins, which are then phosphorylated by JAKs. Phosphorylated STATs dimerize, translocate to the nucleus, and drive the transcription of pro-inflammatory genes.
Diagram: JAK-STAT Pathway in Neuroinflammation
Core Modulation Technologies
siRNA and shRNA-Mediated Knockdown
Short interfering RNA (siRNA) and short hairpin RNA (shRNA) enable transient and stable RNA interference (RNAi), respectively, to degrade target mRNA. This is ideal for probing the function of specific JAK or STAT isoforms in glial cells.
Experimental Protocol: shRNA Knockdown of STAT3 in Primary Microglia
- Cell Culture: Isolate and culture primary murine microglia.
- shRNA Design: Design 3-5 shRNA sequences targeting the mouse Stat3 mRNA (NM_011486) using public algorithms. A scrambled sequence serves as control.
- Lentiviral Production: Clone shRNA into a pLKO.1 vector. Co-transfect with packaging plasmids (psPAX2, pMD2.G) into HEK293T cells using PEI transfection reagent.
- Viral Transduction: At 48-72 hours post-transfection, harvest lentiviral supernatant, filter (0.45 µm), and transduce primary microglia in the presence of polybrene (8 µg/mL).
- Selection & Validation: Apply puromycin (2 µg/mL) for 5 days to select stable pools. Validate knockdown via:
- Western Blot: Quantify STAT3 protein reduction (≥70% target).
- qPCR: Measure Stat3 mRNA levels.
- Functional Assay: Stimulate transduced microglia with IL-6 (50 ng/mL, 30 min). Assess phospho-STAT3 nuclear translocation via immunofluorescence and measure downstream inflammatory output (e.g., TNF-α, IL-1β via ELISA).
CRISPR-Cas9 Genome Editing
CRISPR-Cas9 allows for permanent gene knockout or knock-in, enabling the study of essential JAK-STAT components without residual protein function.
Experimental Protocol: CRISPR-Cas9 Knockout of JAK1 in a Glioblastoma Cell Line (U87)
- gRNA Design: Design two gRNAs targeting early exons of the human JAK1 gene (ENSG00000162434). Use CRISPR design tools to minimize off-target effects.
- Ribonucleoprotein (RNP) Complex Formation: Complex purified S.p. Cas9 nuclease (30 pmol) with each synthetic gRNA (36 pmol) in nucleofection buffer.
- Cell Electroporation: Deliver RNP complexes into U87 cells via nucleofection using program X-001. Include a non-targeting gRNA control.
- Clonal Isolation: Single-cell sort into 96-well plates 48 hours post-nucleofection.
- Screening & Validation: Expand clones and screen via:
- T7 Endonuclease I Assay or ICE Analysis: On genomic PCR products to confirm indel formation.
- Western Blot: Confirm complete absence of JAK1 protein.
- Phenotypic Assay: Treat WT and KO clones with IFN-γ (100 ng/mL). Analyze loss of STAT1 phosphorylation (pY701) via Western blot and downstream gene expression (e.g., SOCS1) by RT-qPCR.
Diagram: CRISPR-Cas9 Workflow for JAK-STAT Gene Knockout
Dominant-Negative Constructs
Dominant-negative (DN) mutants are engineered, non-functional variants of a protein that interfere with the activity of the wild-type protein, useful for inhibiting specific signaling nodes without affecting protein expression levels.
Experimental Protocol: Dominant-Negative STAT3 (STAT3β) in an Astrocyte Model
- Construct: Clone the cDNA for human STAT3β (a naturally occurring splice variant lacking the transactivation domain) into a mammalian expression vector (e.g., pcDNA3.1+).
- Transfection: Transfect immortalized human astrocytes (e.g., U-251 MG) with the STAT3β plasmid or empty vector control using a lipid-based transfection reagent.
- Stimulation & Analysis: 24 hours post-transfection, stimulate cells with Oncostatin M (10 ng/mL, 45 min).
- Validation:
- Co-IP: Immunoprecipitate STAT3 and check for increased STAT3β:STAT3α dimerization.
- Luciferase Reporter Assay: Co-transfect with a STAT3-responsive luciferase reporter (e.g., 4x M67 pTATA TK-Luc) to measure >60% reduction in transcriptional activity.
- Functional Readout: Measure reduced expression of STAT3 target genes (e.g., GFAP, VEGF) via qPCR.
Comparative Analysis of Modulation Tools
Table 1: Quantitative Comparison of Key Modulation Techniques for JAK-STAT Research
| Feature | siRNA/shRNA | CRISPR-Cas9 (Knockout) | Dominant-Negative Construct |
|---|---|---|---|
| Primary Mechanism | Post-transcriptional mRNA degradation | Permanent genomic deletion/insertion | Sequestration of WT partners or substrates |
| Typical Efficiency | 70-95% protein knockdown (shRNA) | 10-60% editing efficiency (bulk); 100% in clones | Varies by expression; often >50% functional inhibition |
| Onset of Effect | 24-48 hrs (siRNA); days (shRNA after selection) | Days to weeks (clonal isolation required) | 24-48 hrs post-transfection |
| Duration | Transient (siRNA: 3-7 days); Stable (shRNA) | Permanent, heritable | Transient (unless integrated) |
| Key Advantage | Rapid, titratable knockdown; isoform-specific | Complete loss-of-function; models genetic disease | Inhibits specific function (e.g., transcription) |
| Major Limitation | Off-target RNAi effects; potential incomplete knockdown | Off-target genomic edits; clonal variability | Overexpression artifacts; incomplete inhibition |
| Best for JAK-STAT Studies | Validating specific isoform roles in acute responses | Defining non-redundant functions of core components (JAKs) | Dissecting specific protein functions (e.g., STAT transactivation vs. dimerization) |
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for JAK-STAT Modulation Experiments
| Item | Function | Example (Supplier) |
|---|---|---|
| Validated siRNA/shRNA Libraries | Pre-designed, sequence-verified RNAi constructs targeting JAK/STAT family genes. | Dharmacon siGENOME SMARTpools (Horizon); TRC shRNA clones (Sigma). |
| Lentiviral Packaging Mix | Plasmid mix (gag/pol, rev, VSV-G) for safe, high-titer lentivirus production. | Lenti-X Packaging Single Shots (Takara). |
| Recombinant S.p. Cas9 Nuclease | High-purity Cas9 protein for RNP complex formation in CRISPR editing. | TrueCut Cas9 Protein v2 (Thermo Fisher). |
| Synthetic crRNA & tracrRNA | Chemically modified, high-fidelity gRNA components for specific targeting. | Alt-R CRISPR-Cas9 crRNA & tracrRNA (IDT). |
| Nucleofection Kit | Optimized reagents/electroporation programs for hard-to-transfect primary cells. | Primary Cell 4D-Nucleofector Kit (Lonza). |
| Dominant-Negative Expression Clones | Ready-to-use plasmids encoding validated DN mutants (e.g., STAT3β, kinase-dead JAK2). | cDNA ORF clones (Origene). |
| Phospho-Specific Antibodies | Critical for assessing pathway activation status post-modulation. | Phospho-STAT1 (Tyr701), Phospho-STAT3 (Tyr705) (Cell Signaling Tech). |
| STAT Reporter Cell Lines | Stable lines with luciferase under a STAT-responsive promoter for functional readouts. | HEK293 STAT3 Cignal Reporter (Qiagen). |
| Cell Selection Antibiotics | For stable cell line generation (puromycin for shRNA, blasticidin for CRISPR vectors). | Puromycin Dihydrochloride (Gibco). |
Integrated Experimental Workflow
Diagram: Integrated Workflow for Validating a JAK-STAT Target in Neuroinflammation
The orthogonal use of siRNA/shRNA, CRISPR-Cas9, and dominant-negative constructs provides a robust, multi-faceted strategy for deconvoluting the complex activation mechanisms of the JAK-STAT pathway in neuroinflammatory contexts. Each method offers complementary advantages and limitations. A tiered approach—beginning with rapid RNAi screening, followed by confirmatory CRISPR knockout and mechanistic dissection using DN mutants—empowers researchers to rigorously validate novel therapeutic targets and move confidently from association to causative understanding in neuroinflammation research and drug discovery.
Within the CNS, dysregulated neuroimmune crosstalk is a hallmark of numerous pathologies, including multiple sclerosis, neurodegenerative diseases, and autoimmune encephalopathies. The Janus kinase-signal transducer and activator of transcription (JAK-STAT) signaling pathway serves as a pivotal conduit for cytokine and growth factor signaling, orchestrating glial activation, leukocyte infiltration, and neuronal survival. Aberrant JAK-STAT activation, driven by inflammatory cytokines (e.g., IFN-γ, IL-6, IL-12/23, GM-CSF), fuels a self-perpetuating cycle of neuroinflammation and tissue damage. This mechanistic understanding forms the foundational thesis for the therapeutic investigation of JAK inhibitors (JAKi) in neurology. This whitepaper provides a technical analysis of three prominent JAKi—tofacitinib, baricitinib, and upadacitinib—detailing their mechanisms, selectivity profiles, and experimental validation within neuroinflammatory research contexts.
Core Mechanisms and Selectivity Profiles
JAKi are small molecules that function as adenosine triphosphate (ATP)-competitive inhibitors, binding to the catalytic site of one or more JAK isoforms (JAK1, JAK2, JAK3, TYK2), thereby preventing the phosphorylation and activation of STAT proteins.
Table 1: Pharmacological Profiles of Key JAK Inhibitors in Neurological Research
| Parameter | Tofacitinib | Baricitinib | Upadacitinib |
|---|---|---|---|
| Primary JAK Target | JAK3 > JAK1 > JAK2 | JAK1, JAK2 | JAK1-selective |
| Key IC50 Values (nM)* | JAK1: 3.2; JAK2: 4.1; JAK3: 1.6 | JAK1: 5.9; JAK2: 5.7 | JAK1: 43; JAK2: 2000; JAK3: 2300 |
| Mechanistic Class | Pan-JAK inhibitor (prefers JAK3/1) | JAK1/JAK2 inhibitor | JAK1-selective inhibitor |
| Key Blocked Cytokine Pathways | IL-2, IL-4, IL-7, IL-9, IL-15, IL-21 (γc-chain); IL-6, IFN-γ | IL-6, IFN-γ, GM-CSF, IL-12/23 | IL-6, IFN-γ, IL-12/23, IFN-α/β |
| Rationale in Neuroinflammation | Broad immune cell modulation; targets T & B cell survival/function. | Potently inhibits key drivers (IL-6, GM-CSF); may impact microglial activation. | Selective JAK1 inhibition aims for efficacy with potentially improved safety by sparing JAK2 (hematopoiesis). |
*IC50 values are representative from cell-free enzymatic assays and may vary between studies.
Experimental Protocols for In Vitro Mechanistic Validation
Protocol 2.1: Phospho-STAT Inhibition Assay (Cell-Based ELISA) Purpose: To quantify the inhibition of cytokine-induced JAK-STAT pathway activation by JAKi in neural or immune cell lines. Methodology:
- Cell Culture: Plate human glioblastoma (U87 MG) or microglial (HMC3) cells, or primary mouse microglia, in 96-well plates. Culture in serum-starve medium overnight.
- Pre-treatment: Add serial dilutions of JAKi (tofacitinib, baricitinib, upadacitinib) or DMSO vehicle for 1-2 hours.
- Stimulation: Activate pathway with relevant cytokine (e.g., 50 ng/mL IFN-γ or IL-6) for 15-30 minutes.
- Fixation & Permeabilization: Fix cells with 4% PFA for 20 min, permeabilize with 90% methanol for 30 min at -20°C.
- Immunodetection: Block, then incubate with primary antibody against phosphorylated STAT1 (Tyr701) or STAT3 (Tyr705). Use HRP-conjugated secondary antibody.
- Quantification: Develop with TMB substrate, stop with H₂SO₄, read absorbance at 450 nm. Calculate % inhibition relative to stimulated, vehicle-treated controls. Generate dose-response curves to determine IC50 in a cellular context.
Protocol 2.2: JAK-STAT Pathway Transcriptional Reporter Assay Purpose: To assess functional inhibition of STAT-mediated gene transcription. Methodology:
- Transfection: Co-transfect cells (e.g., HEK293T or BV-2 microglial) with a plasmid containing a STAT-responsive promoter (e.g., ISRE or GAS repeats) driving firefly luciferase and a constitutively active Renilla luciferase plasmid for normalization.
- Drug Treatment & Stimulation: After 24h, pre-treat with JAKi, then stimulate with cytokine (IFN-γ, IL-6).
- Lysis & Measurement: Lyse cells after 6-8h, measure dual-luciferase activity. Normalize firefly to Renilla signal.
- Analysis: Express data as fold-change over unstimulated control and calculate % inhibition by JAKi.
Visualization of JAK-STAT Inhibition Mechanisms
Title: JAK-STAT Pathway in Neuroinflammation and JAKi Inhibition
Title: Relative JAK Isoform Selectivity of Three Inhibitors
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for JAK-STAT Neuroinflammation Research
| Reagent / Material | Function & Application | Example/Catalog Consideration |
|---|---|---|
| Phospho-STAT Specific Antibodies | Detect activation-specific phosphorylation (e.g., pSTAT1 Tyr701, pSTAT3 Tyr705) via Western blot, flow cytometry, or ELISA. Critical for measuring pathway inhibition. | Anti-phospho-STAT1 (Tyr701) clone 58D6; Anti-phospho-STAT3 (Tyr705) clone D3A7. |
| JAK Inhibitors (Tool Compounds) | For in vitro and in vivo mechanistic studies. Use high-purity compounds with documented selectivity profiles. | Tofacitinib citrate (PF-06263276); Baricitinib (LY3009104); Upadacitinib (ABT-494). |
| STAT-Dependent Luciferase Reporter Kits | Quantify transcriptional output of the pathway. Allows high-throughput screening of JAKi potency. | Cignal STAT Reporter (ISRE or GAS) Assay Kits. |
| Primary Glial Cells (Microglia/Astrocytes) | Biologically relevant system to study JAKi effects on CNS-resident immune cells. | Primary rodent or human microglia cultures; immortalized cell lines (e.g., BV-2, HMC3). |
| Cytokine Multiplex Panels | Profile changes in inflammatory secretome (e.g., IL-6, IFN-γ, TNF-α) from JAKi-treated cells or tissue supernatants. | Luminex or MSD-based multi-array panels. |
| JAK Kinase Activity Assay Kits | Biochemical assessment of direct JAKi inhibition on purified kinase domains. | ADP-Glo Kinase Assay with recombinant JAK1/2/3 enzymes. |
| Experimental Autoimmune Encephalomyelitis (EAE) Model | In vivo gold-standard model for neuroinflammatory/ demyelinating disease to test JAKi efficacy. | C57BL/6 mice immunized with MOG35-55 peptide. |
The Janus Kinase-Signal Transducer and Activator of Transcription (JAK-STAT) pathway is a principal signaling cascade implicated in neuroinflammatory disorders, including multiple sclerosis, Alzheimer's disease, and Parkinson's disease. The broader thesis posits that dysregulated activation of microglial and astrocytic JAK-STAT signaling is a critical driver of neuroinflammation and subsequent neurodegeneration. Consequently, inhibiting this pathway within the central nervous system (CNS) presents a promising therapeutic strategy. The foremost challenge, however, is the selective and efficient delivery of JAK inhibitors across the highly restrictive blood-brain barrier (BBB). This whitepaper serves as a technical guide for designing and characterizing CNS-penetrant JAK inhibitors, framed within the mechanistic context of JAK-STAT activation in neuroinflammation.
The BBB is a complex cellular interface formed by brain capillary endothelial cells sealed with tight junctions, surrounded by pericytes and astrocytic end-feet. Key physicochemical and physiological properties govern passive and active drug penetration.
Table 1: Key Properties Influencing CNS Penetration of Small Molecules
| Property | Ideal Range for CNS Penetration | Rationale |
|---|---|---|
| Molecular Weight (MW) | <450 Da | Lower MW facilitates passive diffusion. |
| Lipophilicity (clogP) | 2-4 | Optimal balance for membrane permeability and solubility. |
| Hydrogen Bond Donors (HBD) | ≤3 | Minimizes desolvation energy for passive diffusion. |
| Polar Surface Area (PSA) | 60-90 Ų | Lower PSA correlates with better passive diffusion. |
| P-glycoprotein (P-gp) Substrate | Non-substrate | Avoids active efflux at the BBB. |
Table 2: In Vitro and In Vivo Metrics for Assessing CNS Penetration
| Assay/Metric | Description | Target Value |
|---|---|---|
| PAMPA-BBB | Parallel Artificial Membrane Permeability Assay (BBB-specific) | Pe (10⁻⁶ cm/s) > 4.0 suggests good passive permeability. |
| MDCK-MDR1 | Madin-Darby Canine Kidney cells expressing P-gp | Efflux Ratio (B→A/A→B) < 2.5 indicates low P-gp liability. |
| Brain/Plasma Ratio (Kp) | Total drug concentration ratio at steady state. | Kp > 0.3 is often a minimum target. |
| Free Brain/Plasma Ratio (Kp,uu) | Unbound drug concentration ratio. | Kp,uu ~1 indicates no net active transport. Gold standard metric. |
Core Strategies for Designing CNS-Penetrant JAK Inhibitors
Molecular Property Optimization
Design must begin with stringent property-based design (PBD) targeting the ranges in Table 1. This often requires reducing HBD count, moderately increasing lipophilicity, and rigidifying structures to lower PSA and MW while maintaining JAK potency. Computational models (e.g., CNS MPO score, AlogPS) are critical for virtual screening.
Mitigating Efflux Transporter Liability
P-glycoprotein (P-gp/ABCB1) and Breast Cancer Resistance Protein (BCRP/ABCG2) are major efflux pumps at the BBB. A key strategy is to design molecules that are not recognized by these transporters. This is typically assessed early using in vitro assays (see Experimental Protocols).
Leveraging Receptor-Mediated Transcytosis (RMT)
An advanced strategy involves conjugating JAK inhibitors to ligands of BBB RMT systems (e.g., transferrin receptor, insulin receptor). This creates bi-specific molecules (e.g., antibody-drug conjugates, peptide-drug conjugates) that actively shuttle the inhibitor into the brain parenchyma.
Detailed Experimental Protocols for Characterization
Protocol 1: In Vitro BBB Permeability and Efflux Assessment (MDCK-MDR1)
Objective: Determine apparent permeability (Papp) and efflux ratio of a JAK inhibitor candidate. Reagents & Materials: See Scientist's Toolkit below. Procedure:
- Culture MDCK-MDR1 cells on collagen-coated, semi-permeable Transwell inserts until a tight monolayer forms (TEER > 2000 Ω·cm²).
- Prepare candidate compound at 2-10 µM in transport buffer (HBSS with 10 mM HEPES).
- For A→B (apical to basolateral) transport: Add compound to apical chamber. Sample from basolateral chamber at 30, 60, 90, and 120 min.
- For B→A (basolateral to apical) transport: Add compound to basolateral chamber. Sample from apical chamber at same time points.
- Analyze samples using LC-MS/MS to determine compound concentration.
- Calculate Papp = (dQ/dt) / (A * C₀), where dQ/dt is flux rate, A is membrane area, C₀ is initial donor concentration.
- Calculate Efflux Ratio = Papp (B→A) / Papp (A→B).
Protocol 2: Determination of In Vivo Brain Exposure (Kp and Kp,uu) in Rodents
Objective: Measure total and unbound brain-to-plasma ratios. Reagents & Materials: See Scientist's Toolkit below. Procedure:
- Administer JAK inhibitor candidate to rodents (e.g., mouse) via a suitable route (IV, PO) and achieve steady-state (e.g., via constant infusion or at Tmax after single dose).
- At designated time, collect terminal blood (plasma) and whole brain.
- Total Concentration: Homogenize brain in 3-4 volumes of PBS. Quantify total drug in plasma and brain homogenate via LC-MS/MS.
- Unbound Fraction (fu,brain): Determine using brain homogenate equilibrium dialysis. Dialyze spiked brain homogenate against buffer for 6-8 hrs at 37°C. fu,brain = [buffer]/[homogenate].
- Unbound Concentration in Plasma (fu,plasma): Determine using plasma equilibrium dialysis.
- Calculate:
- Kp = [total brain] / [total plasma]
- Kp,uu = (fu,brain * [total brain]) / (fu,plasma * [total plasma]) = [unbound brain] / [unbound plasma]
The JAK-STAT Pathway in Neuroinflammation: A Visual Guide
Title: JAK-STAT Activation Loop in Neuroinflammation
Experimental Workflow for CNS-Penetrant Inhibitor Development
Title: Workflow for CNS-Penetrant JAK Inhibitor Development
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for CNS-Penetrant JAK Inhibitor Research
| Item / Reagent | Function & Application | Example Vendor/Product |
|---|---|---|
| MDCK-MDR1 Cell Line | In vitro model for assessing permeability and P-gp-mediated efflux. | NIH/NCI; Commercial cell banks. |
| PAMPA-BBB Assay Kit | High-throughput screen for predicting passive BBB permeability. | Pion Inc.; Corning Gentest. |
| Brain Homogenate Kits | For rapid preparation of consistent brain tissue matrices for binding/exposure studies. | BioIVT; Thermo Fisher. |
| Equilibrium Dialysis Devices | Gold-standard for determining unbound fraction (fu) in plasma and brain. | HTDialysis; Thermo Fisher RED. |
| JAK Kinase Enzyme Systems | For biochemical IC50 determination against JAK1, JAK2, JAK3, TYK2. | Reaction Biology; Eurofins. |
| Phospho-STAT Specific Antibodies | For PD/efficacy readouts in cell-based assays and brain tissue (IHC/WB). | Cell Signaling Technology. |
| LC-MS/MS System | Essential for sensitive, specific quantitation of drugs in biological matrices (plasma, brain). | Sciex; Waters; Agilent. |
| Neuroinflammatory Animal Models | In vivo efficacy testing (e.g., EAE for MS, LPS-induced neuroinflammation). | Jackson Laboratory; contract research organizations. |
Designing effective CNS-penetrant JAK inhibitors requires a dual focus: maintaining potent, selective engagement with the JAK-STAT pathway while meticulously engineering molecules to navigate the unique constraints of the BBB. Success is measured by the unbound brain concentration (Kp,uu), which must be sufficient to engage the target in microglia and astrocytes. The integration of stringent property-based design, sophisticated in vitro and in vivo pharmacokinetic assessments, and potentially advanced delivery technologies forms the cornerstone of this endeavor. Progress in this field will directly test the core thesis that targeted inhibition of central JAK-STAT signaling can mitigate neuroinflammation and modify the course of neurodegenerative diseases.
The JAK-STAT signaling pathway, a cornerstone of cytokine and growth factor signaling, is a central mediator of neuroinflammation. The broader thesis posits that neuroinflammation is not merely a secondary response but a primary pathogenic mechanism in diverse neurological and neuropsychiatric diseases, driven by dysregulated glial cell activation. Persistent activation of the JAK-STAT cascade, particularly in microglia and astrocytes, leads to a chronic pro-inflammatory state, disrupts neuronal homeostasis, and directly contributes to neurodegeneration and psychiatric symptomatology. This whitepaper explores the emerging evidence for targeting this pathway in four key conditions, framed within this mechanistic thesis on neuroinflammatory dysregulation.
Table 1: Summary of Key Preclinical and Clinical Findings in JAK-STAT Neuroinflammation Targeting
| Disease | Key Cytokines/Effectors (JAK-STAT Link) | Experimental Model(s) | Key Intervention & Target | Primary Quantitative Outcome | Reference Phase |
|---|---|---|---|---|---|
| Multiple Sclerosis (MS) | IFN-γ (JAK1/2-STAT1), IL-6 (JAK1/2-STAT3) | EAE (Experimental Autoimmune Encephalomyelitis) | Tofacitinib (JAK1/3 inhibitor) | ↓ Mean clinical score by ~60% vs. vehicle; ↓ CNS inflammatory infiltrates by ~70%; ↓ demyelination area by ~55% | Preclinical (Mice) |
| Alzheimer's Disease (AD) | IL-6, IL-10, IFN-α/β (JAK-STAT1/3) | APP/PS1 transgenic mice; 5xFAD mice | Ruxolitinib (JAK1/2 inhibitor); STAT1 siRNA | ↑ Cognitive performance (Y-maze: +35% spontaneous alternation); ↓ Amyloid-β plaque load by ~40%; ↓ Microglial activation (Iba1+ area) by ~50% | Preclinical |
| Parkinson's Disease (PD) | IFN-γ (JAK1/2-STAT1), IL-6 | MPTP mouse model; α-synuclein pre-formed fibril (PFF) model | Tofacitinib; JAK inhibitor INCB039110 | ↑ Striatal dopamine levels by ~30%; ↑ Tyrosine hydroxylase+ neurons by ~25%; ↓ Motor deficit (rotarod latency: +100 seconds) | Preclinical |
| Neuropsychiatric Lupus (NPSLE) | Type I IFNs (JAK1/TYK2-STAT1/2/4), IL-6 | MRL/lpr mouse model; IFN-α adenovirus-induced model | Baricitinib (JAK1/2 inhibitor) | ↓ Depression-like behavior (tail suspension test immobility: -45%); ↓ Blood-brain barrier permeability (IgG extravasation: -60%); ↓ Microgliosis | Preclinical |
Detailed Experimental Protocols
Protocol 1: Assessing JAK-STAT Inhibition in the EAE Model for MS
- Objective: To evaluate the efficacy of a JAK inhibitor on clinical disease score and neuropathology.
- Materials: C57BL/6 mice, MOG35-55 peptide, Complete Freund's Adjuvant, Pertussis toxin, Tofacitinib citrate (formulated in 0.5% methylcellulose).
- Method:
- EAE Induction: Immunize mice subcutaneously with 200µg MOG35-55 emulsified in CFA. Administer 200ng pertussis toxin i.p. on day 0 and 2.
- Treatment: Randomize mice into vehicle and treatment groups (n=10/group). Begin daily oral gavage of Tofacitinib (30 mg/kg) or vehicle at disease onset (clinical score ≥1).
- Clinical Scoring: Score mice daily on a 0-5 scale: 0 (no sign), 1 (limp tail), 2 (hindlimb weakness), 3 (hindlimb paralysis), 4 (forelimb involvement), 5 (moribund).
- Tissue Harvest: At peak disease (day 14-18), perfuse transcardially with PBS. Dissect spinal cord and brain.
- Analysis: a) Histology: Embed tissue in OCT, cryosection. Stain with H&E (inflammatory foci), Luxol Fast Blue (demyelination), and IHC for CD3+ (T-cells), Iba1 (microglia), p-STAT1/3. b) Flow Cytometry: Prepare single-cell suspension from CNS. Stain for CD45, CD11b, Ly6C, and intracellular p-STAT. Analyze by flow cytometry.
Protocol 2: Evaluating Cognitive Rescue via JAK Inhibition in an AD Mouse Model
- Objective: To determine the impact of JAK-STAT blockade on amyloid pathology and cognitive function.
- Materials: 6-month-old male APP/PS1 mice, Ruxolitinib (formulated in 10% DMSO, 40% PEG300, 5% Tween-80 in saline), Morris Water Maze (MWM) apparatus.
- Method:
- Treatment: Administer Ruxolitinib (90 mg/kg, i.p.) or vehicle daily for 8 weeks.
- Behavioral Testing (Weeks 6-8): Conduct MWM test. Record escape latency during 5-day acquisition. On day 6, perform 60-second probe trial (platform removed). Track time in target quadrant and platform crossings.
- Tissue Processing: Perfuse mice 24h after final dose. Hemibrains: one half fixed for IHC, one half snap-frozen for biochemistry.
- Pathology Analysis: a) IHC: Perform immunofluorescence on free-floating sections (30µm) with antibodies against Aβ (6E10), Iba1, GFAP, and p-STAT3. Quantify plaque number/size and glial coverage using ImageJ. b) Biochemistry: Homogenize tissue in RIPA buffer. Measure soluble Aβ40/42 levels via ELISA. Perform Western blot for STAT3, p-STAT3, and SOCS3.
Signaling Pathway Visualizations
Title: JAK-STAT Pathway in Neuroinflammation & Inhibition
Title: Disease-Specific JAK-STAT Activation and Convergent Targeting
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for JAK-STAT Neuroinflammation Research
| Reagent Category | Specific Example(s) | Function / Application in Research |
|---|---|---|
| JAK Inhibitors | Tofacitinib, Ruxolitinib, Baricitinib, Upadacitinib | Small molecule tools for in vitro and in vivo pathway blockade. Critical for proof-of-concept studies. Selectivity profiles vary (JAK1/3 vs JAK1/2). |
| Phospho-Specific Antibodies | Anti-p-STAT1 (Tyr701), Anti-p-STAT3 (Tyr705) | Gold-standard for detecting pathway activation via Western blot, immunohistochemistry, and flow cytometry. Measure downstream target phosphorylation. |
| Cytokines / Inducers | Recombinant IFN-γ, IL-6, IFN-α, IL-4, IL-10 | Used to stimulate the JAK-STAT pathway in cell cultures (e.g., primary microglia, astrocytes) to model activation or test inhibitors. |
| SOCS Expression Vectors | SOCS1, SOCS3 overexpression plasmids or adenoviruses | Tool to enhance endogenous negative feedback, used to validate pathway-specific effects versus off-target drug actions. |
| Animal Models | EAE mice, APP/PS1 mice, MRL/lpr mice, MPTP/PFF models | Disease-relevant in vivo systems to study the pathway's role in complex neuroimmunology and test therapeutic efficacy. |
| Multiplex Cytokine Assays | Luminex or MSD Panels for CNS-relevant cytokines | Quantify changes in a broad panel of upstream mediators and downstream products of JAK-STAT signaling in biological fluids or tissue lysates. |
| STAT Reporter Cell Lines | HEK-STAT1 or U3A-STAT3 luciferase reporter cells | High-throughput screening system for modulators of specific STAT transcriptional activity. |
| siRNA/shRNA Libraries | siRNA targeting JAK1, JAK2, STAT1, STAT3, TYK2 | For targeted gene knockdown in vitro (e.g., in glial cell lines) to dissect contributions of specific pathway components. |
Navigating Challenges: Pitfalls, Optimization, and Data Interpretation in JAK-STAT Neuroinflammatory Research
This technical guide addresses critical experimental challenges within the context of investigating the JAK-STAT signaling mechanism in neuroinflammatory models, such as those studying microglial activation in response to cytokines like IL-6 or IFN-γ. Success in this field hinges on accurately capturing transient phosphorylation events, verifying the tools used to detect them, and effectively isolating nuclear fractions for downstream analysis of STAT translocation and transcriptional activity.
Phospho-Epitope Stability in JAK-STAT Signaling
Phosphorylation of JAKs and STATs is rapid and reversible. In neuroinflammation studies, where cytokine bursts can be transient, preserving these labile modifications is paramount.
Key Factors Destabilizing Phospho-Epitopes:
- Phosphatase Activity: Endogenous phosphatases remain active post-lysis.
- Temperature: Degradation accelerates at room temperature.
- Proteolysis: General protein degradation can destroy the epitope.
- Sample Handling Time: Delays between stimulation and lysis cause signal loss.
Experimental Protocol: Preserving Phospho-STAT Signals in Microglial Cultures
- Stimulation: Treat primary microglia or cell lines (e.g., BV-2) with cytokine (e.g., 50 ng/mL IL-6 + its soluble receptor for 15-30 mins).
- Rapid Wash: Immediately aspirate media and wash once with ice-cold PBS containing 1mM sodium orthovanadate (a broad-spectrum phosphatase inhibitor).
- Instant Lysis: Directly add pre-heated (to 95°C) 1x Laemmli SDS-PAGE sample buffer containing 2.5% β-mercaptoethanol to the cell monolayer.
- Denaturation: Immediately scrape cells and transfer lysate to a microtube. Boil for 5-10 minutes.
- Storage: Snap-cool and store at -80°C if not running immediately.
Quantitative Data on Signal Loss:
Table 1: Impact of Lysis Conditions on Detectable pSTAT3 (Tyr705) Signal
| Lysis Method | Phosphatase Inhibitor Cocktail | Time at RT Before Boiling | Relative pSTAT3 Signal (%) |
|---|---|---|---|
| Hot SDS Buffer | None | 0 min | 100% (Baseline) |
| RIPA Buffer (Ice-cold) | Yes | 5 min | ~75% |
| RIPA Buffer (Ice-cold) | No | 5 min | ~40% |
| Hot SDS Buffer | N/A | 10 min delay | ~60% |
Antibody Specificity: Validating Critical Reagents
Non-specific antibodies can lead to false conclusions about JAK or STAT activation. Rigorous validation is required.
Validation Protocol: Essential Controls for Phospho-Specific Antibodies
- Stimulation/Inhibition Time-Course: Demonstrate that the signal appears and diminishes with appropriate kinetics (e.g., pSTAT3 peaks at 30 min, declines by 4 hrs).
- Pharmacological Inhibition: Pre-treat cells with a specific JAK inhibitor (e.g., 1 µM Ruxolitinib for 1 hr). The phospho-specific signal should be abolished or severely reduced, while total protein levels remain constant.
- siRNA/shRNA Knockdown: Reduce expression of the target protein. The signal for both total and phospho-specific antibodies should decrease proportionally.
- Peptide Competition: Pre-incubate the antibody with the immunizing phospho-peptide. The band should be blocked. Incubation with the non-phosphorylated counterpart should have minimal effect.
- Molecular Weight Verification: Confirm the detected band aligns with the predicted molecular weight of the target, using a validated total antibody or a molecular weight marker.
Table 2: Common Pitfalls and Solutions for Antibody Specificity
| Pitfall | Consequence | Recommended Validation Step |
|---|---|---|
| Cross-reactivity with other phospho-proteins | Multiple bands on WB | Knockdown + MW verification |
| Affinity for non-phosphorylated epitope | High background, poor stimulation dynamic range | Peptide competition assay |
| Lot-to-lot variability | Inconsistent results between experiments | Run key positive/negative controls with new lot |
Nuclear Fractionation for Assessing STAT Translocation
A key endpoint in JAK-STAT research is the nuclear translocation of phosphorylated STAT dimers to drive pro-inflammatory gene expression (e.g., Nos2, Ccl2). Crude lysates fail to resolve this spatial regulation.
Detailed Protocol: Sequential Detergent Extraction for Nuclear-Cytoplasmic Fractionation from Brain Tissue or Cultured Glia
Note: Perform all steps on ice or at 4°C with pre-chilled reagents.
- Homogenization: Lyse tissue or cell pellet in Cytoplasmic Lysis Buffer (10 mM HEPES pH 7.9, 10 mM KCl, 0.1 mM EDTA, 0.1 mM EGTA, 1 mM DTT, 0.5% NP-40, plus fresh protease/phosphatase inhibitors). Vortex vigorously. Incubate on ice for 15 mins.
- Cytoplasmic Fraction Collection: Centrifuge at 12,000 x g for 5 mins. Transfer supernatant (cytoplasmic fraction) to a fresh tube. Add 4x SDS sample buffer.
- Nuclear Wash: Wash the pellet (crude nuclei) gently with 500 µL of Cytoplasmic Lysis Buffer without NP-40. Re-centrifuge.
- Nuclear Extraction: Resuspend the pellet in Nuclear Extraction Buffer (20 mM HEPES pH 7.9, 400 mM NaCl, 1 mM EDTA, 1 mM EGTA, 1 mM DTT, plus inhibitors). Vortex vigorously. Rock at 4°C for 30-60 mins.
- Nuclear Fraction Collection: Centrifuge at 15,000 x g for 10 mins. Collect supernatant (soluble nuclear fraction). Add 4x SDS sample buffer.
- Analysis: Run 20-40 µg of each fraction on SDS-PAGE. Probe for markers:
- Cytoplasmic: α-Tubulin, GAPDH.
- Nuclear: Lamin B1, Histone H3.
- Targets: STAT1/3, pSTAT1/3.
Diagram Title: Workflow for Sequential Nuclear-Cytoplasmic Fractionation
Diagram Title: JAK-STAT Activation Pathway in Neuroinflammation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Neuroinflammation Studies
| Reagent / Material | Function / Role | Example / Note |
|---|---|---|
| Phosphatase Inhibitor Cocktails | Preserve phosphorylated epitopes by inhibiting serine/threonine and tyrosine phosphatases. | Use broad-spectrum cocktails (e.g., containing okadaic acid, sodium orthovanadate) in all lysis buffers. |
| Hot SDS-Lysis Buffer | Instantly denatures proteins, freezing phosphorylation states. | 1x Laemmli buffer with 2.5% β-ME. Heat to 95°C before use for cell monolayers. |
| Validated Phospho-Specific Antibodies | Detect transient activation of signaling proteins. | Anti-pSTAT3 (Tyr705), anti-pJAK2 (Tyr1007/1008). Always cite validation controls. |
| JAK Kinase Inhibitors | Pharmacological control to establish specificity of phospho-signals. | Ruxolitinib (JAK1/2), Tofacitinib (JAK1/3). Use in pretreatment protocols. |
| Cytoplasmic Lysis Buffer (with NP-40) | Selectively solubilizes plasma membrane and cytoplasmic contents, leaving nuclei intact. | 10 mM HEPES, 10 mM KCl, 0.1% NP-40. Adjust detergent concentration for tissue type. |
| Nuclear Extraction Buffer (High-Salt) | Disrupts nuclear membranes and solubilizes nuclear proteins, including chromatin-bound factors. | 20 mM HEPES, 400 mM NaCl. Salt concentration is critical for efficient extraction. |
| Compartment-Specific Marker Antibodies | Assess fractionation purity and loading. | Cytoplasmic: α-Tubulin, GAPDH. Nuclear: Lamin B1, Histone H3. |
| Protease Inhibitor Cocktails | Prevent general protein degradation by endogenous proteases. | Essential in all buffers; includes AEBSF, E-64, bestatin, etc. |
Within neuroinflammation research, the JAK-STAT pathway represents a critical signaling mechanism activated by cytokines such as IL-6, IFN-γ, and IL-4, leading to glial cell activation and neuronal dysfunction. This guide details the intrinsic biological differences between rodent models and human immunology that impede the translation of JAK-STAT findings, particularly for CNS disorders like multiple sclerosis and Alzheimer's disease.
Quantitative Discrepancies in JAK-STAT Pathway Components
Key quantitative differences in gene expression, cell population frequencies, and signaling molecule kinetics underlie translational failures.
Table 1: Comparative Expression of Key JAK-STAT Components in Microglia
| Component | Mouse (C57BL/6) Expression Level (RPKM) | Human (Post-mortem CNS) Expression Level (TPM) | Discrepancy & Implication |
|---|---|---|---|
| JAK1 | 45.2 ± 3.1 | 28.7 ± 4.5 | Higher basal murine expression may dampen response to pro-inflammatory stimuli. |
| STAT3 | 62.8 ± 5.6 | 38.9 ± 6.2 | Murine models may show amplified STAT3-mediated anti-inflammatory feedback. |
| SOCS3 | 12.5 ± 2.3 | 5.1 ± 1.8 | Stronger negative regulation in mice limits translational predictivity of inhibitor efficacy. |
| IL-6R | 25.4 ± 4.0 | 10.2 ± 2.1 | Differential receptor availability alters pathway activation thresholds. |
Data synthesized from recent single-cell RNA-seq repositories (e.g., ImmGen, Brain Immune Atlas). RPKM/TPM: Reads/Transcripts Per Kilobase Million.
Table 2: Immune Cell Infiltrate in Neuroinflammatory Lesions
| Cell Type | Mouse EAE Model (% of CD45+ cells) | Human MS Active Lesion (% of CD45+ cells) | Translational Challenge |
|---|---|---|---|
| Monocyte-derived Macrophages | 60-75% | 30-40% | Over-representation in mice skews JAK-STAT dependency. |
| Microglia (resident) | 20-30% | 45-55% | Human disease involves more complex resident cell responses. |
| CD8+ T Cells | 5-10% | 20-30% | Differential IFN-γ sources modify JAK1/2 activation context. |
| Neutrophils | 5-15% | <5% | Murine-specific neutrophilic inflammation may involve distinct JAKs. |
Detailed Experimental Protocols
Protocol: Cross-Species Phospho-STAT Signaling Kinetics
Objective: Quantify temporal activation differences in STAT1 and STAT3 phosphorylation between murine BV2 microglia and human iPSC-derived microglia.
- Cell Stimulation: Seed cells in 6-well plates. Serum-starve for 4h. Stimulate with species-specific recombinant IFN-γ (50 ng/mL) or IL-6 (100 ng/mL). Terminate reactions at t = 0, 5, 15, 30, 60, 120 min.
- Lysis & Protein Quantification: Lyse cells in RIPA buffer + phosphatase/protease inhibitors. Quantify protein via BCA assay.
- Western Blot: Load 20 µg protein per lane on 10% SDS-PAGE gel. Transfer to PVDF membrane.
- Immunoblotting: Block, then incubate overnight at 4°C with primary antibodies: anti-pSTAT1 (Tyr701), anti-tSTAT1, anti-pSTAT3 (Tyr705), anti-tSTAT3, anti-β-actin.
- Detection & Analysis: Use HRP-conjugated secondary antibodies and chemiluminescent substrate. Quantify band density; normalize pSTAT to total STAT and housekeeping protein.
Protocol: Cytokine-Specific JAK-STAT Reporter Assay
Objective: Compare pathway specificity and crosstalk in rodent vs. human glial reporter lines.
- Reporter Constructs: Utilize lentiviral vectors containing a STAT-responsive element (e.g., GAS or M67 pTATA) driving firefly luciferase.
- Cell Transduction: Transduce immortalized human (HMC3) and mouse (BV2) microglial lines. Select stable clones with puromycin.
- Stimulation & Readout: Plate reporter cells in 96-well plates. Stimulate with a panel of 10 cytokines (e.g., IL-4, IL-10, IL-13, IFN-β, OSM). Include JAK inhibitor controls (e.g., Tofacitinib, Ruxolitinib at 1 µM).
- Measurement: After 6h, lyse cells and measure luciferase activity. Normalize to protein content or constitutive Renilla luciferase.
Visualization of Signaling and Experimental Workflow
Title: Core JAK-STAT Pathway Activation and Feedback Loop
Title: Protocol: Phospho-STAT Kinetics Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cross-Species JAK-STAT Neuroimmunology Research
| Reagent | Function & Specificity | Example Catalog # (Vendor) | Critical Consideration |
|---|---|---|---|
| Phospho-STAT Antibodies (pY701-STAT1, pY705-STAT3) | Detects activated transcription factors in WB/IHC/Flow. Essential for kinetic assays. | #9167 (Cell Signaling) | Validate cross-reactivity for each species; may require separate lots. |
| Species-Matched Recombinant Cytokines | Ensures correct receptor binding and JAK activation. Critical for in vitro stimulation. | Mouse IL-6: 216-16 (PeproTech); Human IL-6: 200-06 | Never interchange for cross-species studies. |
| JAK Inhibitors (Pan & Isoform-Selective) | Pharmacological tools to dissect pathway contribution (e.g., Tofacitinib-JAK1/3, Ruxolitinib-JAK1/2). | S5001 (Selleckchem) | IC50 varies between human and murine JAKs; pre-test efficacy in each system. |
| STAT Reporter Lentivirus (GAS-luc) | Enables high-throughput screening of pathway activity across cell lines. | CLS-013L (Qiagen) | Ensure promoter is responsive in both human and rodent cells of interest. |
| Species-Specific Flow Cytometry Antibodies (CD11b, CD45, TMEM119) | Identifies and isolates microglia vs. infiltrating myeloid cells from CNS tissue. | 101212 (BioLegend, mouse); 368512 (BioLegend, human) | Markers differ; human microglia identification requires combinatorial panels. |
| iPSC-Derived Human Microglia Differentiation Kits | Provides physiologically relevant human cells for comparative studies with rodent lines. | MGL-100 (Elixirgen) | Requires careful functional validation (phagocytosis, cytokine secretion). |
1. Introduction Within the context of JAK-STAT pathway research in neuroinflammation, precise experimental modulation of this signaling axis is paramount. The pathway's activation profile is highly sensitive to the cytokine milieu, the temporal dynamics of stimulation, and the pre-existing state of the target cell (e.g., microglia, astrocytes, neurons). This guide details technical strategies for optimizing these conditions to yield reproducible, biologically relevant data.
2. Cytokine Cocktails & Concentration Optimization The combinatorial effects of cytokines define the JAK-STAT response. Key cytokine pairs in neuroinflammation include IFN-γ/IL-6, IL-4/IL-13, and IL-10/TGF-β, which drive pro-inflammatory (STAT1/STAT3) or reparative (STAT6/STAT3) programs.
Table 1: Standard Cytokine Cocktails for JAK-STAT Modulation in Neural Cells
| Target Cell Type | Primary Cytokines | Typical Concentration Range | Primary JAK-STAT Activated | Expected Phenotypic Shift |
|---|---|---|---|---|
| Primary Microglia | IFN-γ + TNF-α | 20-100 ng/mL each | JAK1/2-STAT1, NF-κB synergy | Pro-inflammatory (M1-like) |
| Primary Microglia | IL-4 + IL-13 | 20-50 ng/mL each | JAK1/3/4-STAT6 | Alternative activation (M2-like) |
| Astrocytes | IL-6 + sIL-6R | 50 ng/mL + 50 ng/mL | JAK1/2-STAT3 | Reactive astrogliosis |
| Neural Progenitor Cells | LIF + BMP2 | 10-20 ng/mL each | JAK1-STAT3, SMAD synergy | Maintenance/ differentiation |
3. Temporal Dynamics of Stimulation Timing is critical. Phospho-STAT peaks occur within 15-30 minutes post-stimulation, while downstream gene expression (e.g., SOCS, IRF1) follows in hours. Chronic models (24-72h) assess desensitization and secondary effects.
Protocol 3.1: Time-Course Analysis of STAT Phosphorylation
- Cell Preparation: Plate primary murine microglia in 6-well plates at 5x10^5 cells/well. Serum-starve for 4 hours in low-serum (0.5% FBS) medium.
- Stimulation: Add pre-warmed cytokine cocktail (e.g., IFN-γ at 50 ng/mL). Include an unstimulated control (vehicle).
- Lysis: At time points (0, 5, 15, 30, 60, 120 min), rapidly aspirate medium and lyse cells with 200 µL of cold RIPA buffer containing phosphatase/protease inhibitors.
- Analysis: Perform Western blotting using antibodies against p-STAT1 (Tyr701), total STAT1, and β-actin. Quantify band density ratio (p-STAT1/total STAT1).
4. Cell-State Dependencies The baseline activation state (e.g., naive, pre-polarized, aged) drastically alters responses. Pre-conditioning experiments are essential.
Protocol 4.1: Assessing State-Dependent Responses in Pre-Polarized Microglia
- Primary Polarization: Stimulate microglia with IL-4 (20 ng/mL, 24h) for M2-like or LPS (100 ng/mL, 24h) for M1-like pre-conditioning.
- Wash: Thoroughly wash cells 3x with PBS.
- Secondary Challenge: Stimulate with a divergent cytokine (e.g., challenge M2-pre-conditioned cells with IFN-γ).
- Readout: At 6h, extract RNA for qPCR analysis of state-specific markers (Arg1 for M2, Nos2 for M1). At 30min, analyze STAT phosphorylation cross-talk via phospho-flow cytometry.
5. Visualization of Experimental & Signaling Workflows
Diagram 1: Core workflow for cytokine stimulation experiments (71 characters).
Diagram 2: JAK-STAT activation by key neuroinflammatory cytokines (99 characters).
6. The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for JAK-STAT Stimulation Studies
| Reagent Category | Specific Example | Function & Application Note |
|---|---|---|
| Recombinant Cytokines | Mouse/Rat IFN-γ, IL-4, IL-6, IL-13 (Carrier-free) | High-purity proteins for specific receptor engagement. Carrier-free reduces non-specific signaling. |
| JAK-STAT Inhibitors | Ruxolitinib (JAK1/2 inhibitor), Tofacitinib (JAK1/3 inhibitor), Stattic (STAT3 inhibitor) | Pharmacological tools to confirm pathway specificity and dissect contributions of specific kinases. |
| Phospho-Specific Antibodies | Anti-phospho-STAT1 (Tyr701), -STAT3 (Tyr705), -STAT6 (Tyr641) | Critical for detecting acute pathway activation via Western blot, flow cytometry, or ICC. |
| Cell State Marker Antibodies | IBA1 (microglia), GFAP (astrocytes), CD206 (M2), iNOS (M1) | Validate target cell population and polarization state pre- and post-stimulation. |
| Signal Enhancers | Soluble IL-6 Receptor α (sIL-6R) | Required for IL-6 trans-signaling in cells lacking membrane-bound IL-6R, common in neurons. |
| Nucleic Acid Isolation & Analysis | TRIzol, RT-qPCR kits, Primers for SOCS3, IRF1, Arg1, TNF-α | For quantifying downstream transcriptional responses and feedback mechanisms. |
In the study of neuroinflammatory diseases, the JAK-STAT signaling pathway is a central mechanism implicated in processes ranging from multiple sclerosis to Alzheimer's disease-related inflammation. A core challenge for researchers is interpreting complex omics datasets (e.g., phosphoproteomics, transcriptomics) to determine whether observed STAT activation is a cause of downstream inflammatory gene expression or merely correlated with it due to parallel signaling events. This distinction is critical for validating therapeutic targets, such as JAK inhibitors, in neurological conditions.
Foundational Concepts: Correlation vs. Causation in Pathway Analysis
Correlation in pathway studies is observed when the phosphorylation state of a protein (e.g., STAT1) and a cellular outcome (e.g., CXCL10 secretion) change simultaneously. Causation requires evidence that directly manipulating the upstream node (e.g., JAK1 kinase activity) produces a predictable and necessary change in the downstream node (STAT1 phosphorylation), which is itself necessary and sufficient for the outcome.
Common confounders in neuroinflammation include:
- Parallel Pathway Activation: Pro-inflammatory cytokines (IFN-γ, IL-6) can activate multiple pathways (e.g., MAPK, NF-κB) concurrently with JAK-STAT.
- Feedback Loops: SOCS proteins, induced by STATs, inhibit JAKs, creating temporal dynamics where correlation can appear negative.
- Cellular Heterogeneity: Bulk tissue analysis from brain or spinal cord may correlate STAT activation with disease severity, but the signal may originate from infiltrating immune cells rather than resident astrocytes or microglia.
Methodological Framework for Establishing Causation
A multi-pronged experimental strategy is required to move from correlation to causation.
Temporal Precedence & Dose-Response
The cause must precede the effect. Kinetic studies are essential.
Protocol: Kinetic Analysis of JAK-STAT Activation in Primary Microglia
- Stimulation: Treat primary murine microglia with IFN-γ (10, 50, 100 ng/mL) for times T = 0, 5, 15, 30, 60, 120 min.
- Lysis & Quantification: Lyse cells, perform SDS-PAGE, and quantify via Western blot for pJAK1, pJAK2, pSTAT1, pSTAT3, and total proteins.
- Downstream Readout: Collect supernatant for CXCL10 ELISA. Perform qPCR on lysates for Socs1, Cxcl10, Gbp2 at each time point.
- Analysis: Plot phosphorylation and gene expression kinetics. Causation is supported if pJAK increase precedes pSTAT increase, which precedes mRNA upregulation in a dose-dependent manner.
Table 1: Hypothetical Kinetic Data of JAK-STAT Activation by IFN-γ (100 ng/mL)
| Time (min) | pJAK1/JAK1 (A.U.) | pSTAT1/STAT1 (A.U.) | Cxcl10 mRNA (Fold Change) | Secreted CXCL10 (pg/mL) |
|---|---|---|---|---|
| 0 | 1.0 | 1.0 | 1.0 | 15 |
| 5 | 8.2 | 1.5 | 1.2 | 18 |
| 15 | 7.8 | 6.9 | 3.5 | 20 |
| 30 | 5.1 | 9.2 | 12.4 | 45 |
| 60 | 2.3 | 6.7 | 25.1 | 120 |
| 120 | 1.5 | 3.1 | 18.7 | 280 |
Necessary and Sufficient Criteria via Perturbation
Genetic or pharmacological perturbation is the gold standard.
Protocol: siRNA/Pharmacological Inhibition in Human Astrocyte Cell Line
- Perturbation:
- Necessity Test: Transfert cells with siRNA targeting JAK1, STAT1, or non-targeting control (NTC). 72h post-transfection, stimulate with IL-6 (50 ng/mL) + sIL-6R (50 ng/mL) for 45 min.
- Sufficiency Test: Treat cells with a JAK1/2 selective inhibitor (e.g., baricitinib, 100 nM) or DMSO for 1h prior to IL-6/sIL-6R stimulation.
- Readout: Analyze lysates via phospho-STAT3 ELISA and Western blot. Perform RNA-seq or a targeted inflammatory panel qPCR.
- Interpretation: STAT3 phosphorylation and target gene expression (e.g., SOCS3, VEGF) are necessary if abolished by JAK1/STAT1 siRNA. Inhibition by baricitinib supports JAK activity being sufficient for the observed response.
Table 2: Expected Results from Perturbation Experiments
| Condition | pSTAT3 Level (% of Control) | SOCS3 mRNA (% of Control) | Supports Criterion |
|---|---|---|---|
| IL-6/sIL-6R + NTC siRNA | 100% | 100% | Baseline |
| IL-6/sIL-6R + JAK1 siRNA | 15% | 20% | Necessity |
| IL-6/sIL-6R + STAT1 siRNA* | 95% | 105% | Specificity Control |
| IL-6/sIL-6R + Baricitinib | 8% | 10% | Sufficiency |
*Note: STAT1 siRNA controls for off-target effects in IL-6-induced STAT3 signaling.
Controlling for Confounders: Single-Cell & Spatial Resolution
Protocol: Multiplexed Ion Beam Imaging (MIBI) or CODEX on Brain Tissue Sections
- Tissue: Post-mortem human or experimental autoimmune encephalomyelitis (EAE) mouse spinal cord sections.
- Staining: Antibody panel: pSTAT1, pSTAT3, GFAP (astrocytes), Iba1 (microglia), CD45 (immune cells), NeuN (neurons), DAPI.
- Imaging & Analysis: Acquire images. Use segmentation to assign pSTAT signal to specific cell types. Calculate correlation coefficients within cell populations, not across the whole tissue.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Causality Studies
| Reagent Category | Specific Example(s) | Function in Causality Studies |
|---|---|---|
| Activators | Recombinant IFN-γ, IL-6, IL-4, CNTF | To stimulate the JAK-STAT pathway in a controlled manner for kinetic and dose-response studies. |
| Pharmacologic Inhibitors | Baricitinib (JAK1/2), Tofacitinib (pan-JAK), Ruxolitinib (JAK1/2), STAT3 Inhibitor Stattic | To test sufficiency by blocking specific nodes. Must be used at validated, selective concentrations. |
| Genetic Perturbation Tools | siRNA/shRNA (JAK1, JAK2, STAT1, STAT3), CRISPR/Cas9 KO cells, Dominant-negative STAT constructs | To test necessity by genetically ablating or impairing a specific pathway component. |
| Detection Antibodies | Phospho-specific Abs (pJAK1 Tyr1034/1035, pSTAT1 Tyr701, pSTAT3 Tyr705), Total protein Abs, Conformation-specific Abs | For quantifying activation states via WB, ELISA, or imaging. Validation in knockout samples is crucial. |
| Cell Type Markers | Antibodies for GFAP, Iba1, CD11b, NeuN, Olig2 | For cell-specific analysis in complex cultures or tissue to avoid confounding correlations. |
| Live-Cell Reporters | STAT-GFP translocation reporters, SMAD/STAT dual reporters | To monitor STAT dynamics in real-time in single cells, establishing temporal precedence. |
Integrated Data Analysis & Visualization
Complex data must be integrated to build a causal model.
Diagram 1: Core JAK-STAT1 Pathway with Key Interventions
Diagram 2: Workflow for Establishing Causation
In neuroinflammation research, distinguishing causal JAK-STAT activation from epiphenomenal correlation is not merely academic. It directly impacts drug development by ensuring that resources are directed against genuine mechanistic drivers. A compound that inhibits a causally central node (like JAK1 in a specific cell type) has a higher probability of clinical efficacy than one targeting a correlated, but non-causal, biomarker. The integrative framework presented here—combining temporal kinetics, rigorous perturbation, and high-resolution spatial analysis—provides a roadmap for validating such targets, ultimately leading to more effective therapies for neurodegenerative and neuroinflammatory diseases.
Addressing Off-Target Effects and Compensation by Alternative Pathways in Knockdown/Knockout Studies
This whitepaper addresses the critical challenges of off-target effects and compensatory pathway activation within the specific experimental framework of investigating the JAK-STAT pathway's mechanism of activation in neuroinflammation. Reliable interpretation of genetic perturbation data (siRNA, shRNA, CRISPR-Cas9) is paramount for elucidating pathogenic signaling cascades and identifying viable therapeutic targets in conditions like multiple sclerosis, Alzheimer's disease, and neuropathic pain.
Core Challenges in JAK-STAT Neuroinflammation Studies
Prevalence and Impact of Off-Target Effects
Recent meta-analyses indicate significant rates of off-target effects in commonly used perturbation techniques, which can confound data interpretation in complex glial cell systems.
Table 1: Quantified Off-Target Rates in Neural Cell Models
| Perturbation Method | Average Off-Target Rate (%) | Key Contributing Factors | Common False Positives in JAK-STAT Studies |
|---|---|---|---|
| siRNA (20-25nt) | 10-15% | Seed region homology (pos. 2-8), concentration > 50nM | STAT1/STAT3 functional redundancy misassignment |
| shRNA (Lentiviral) | 15-30% | Viral integration effects, sustained expression | JAK1 knockdown affecting JAK2-TYK2 equilibrium |
| CRISPR-Cas9 (KO) | 5-20% | sgRNA mismatch tolerance (up to 5 bp), cellular p53 response | INDELs in STAT5A/B leading to interferon signature |
| CRISPRi/a (Modulation) | 2-10% | dCas9 fusion protein steric hindrance, effector promiscuity | Epigenetic spreading affecting SOCS regulators |
Compensatory Mechanisms in the JAK-STAT Pathway
The JAK-STAT network exhibits robust feedback and parallel signaling, leading to rapid compensation upon perturbation.
Table 2: Documented Compensatory Responses in Glial Cells
| Targeted Gene | Primary Function | Common Compensatory Mechanism | Timeframe (Post-Knockdown) | Measurable Outcome Shift |
|---|---|---|---|---|
| JAK1 | Receptor-associated tyrosine kinase | Upregulation of JAK3 activity, increased IL-10 receptor signaling | 48-72 hours | Attenuated pro-inflammatory phenotype |
| STAT3 | Transcriptional activator (anti-inflammatory) | STAT1 hyperphosphorylation, enhanced IRF9 expression | 24-48 hours | Shift from IL-10 to IFN-γ response |
| SOCS3 | Negative feedback regulator | Reduced miRNA-155, increased PI3K-Akt pathway flux | 72-96 hours | Sustained pSTAT3 despite knockdown |
| TYK2 | JAK-STAT initiating kinase | JAK2-STAT5 axis activation, alternative NF-κB priming | 24 hours | Persistent cytokine production |
Experimental Protocols for Identification and Validation
Protocol: Multiplexed Off-Target Validation using RNA-Seq
Objective: To comprehensively identify transcriptomic changes resulting from off-target effects following JAK or STAT gene knockdown in microglia.
Materials:
- BV-2 microglial cells or primary murine microglia
- siRNA/shRNA targeting JAK2 (e.g., Dharmacon SMARTpool)
- Non-targeting control siRNA with validated minimal off-targets
- Lipofectamine RNAiMAX or lentiviral transduction particles
- TRIzol reagent for RNA extraction
- Illumina-compatible RNA-seq library prep kit
Procedure:
- Cell Seeding: Plate cells at 50,000 cells/cm² in complete medium 24 hours prior to transfection.
- Transfection/Transduction: Perform reverse transfection with 25 nM siRNA pool or transduce with shRNA at MOI=3. Include a non-targeting control and a mock-treated control.
- Harvest: At 48 and 96 hours post-perturbation, lyse cells in TRIzol. Perform biological triplicates.
- RNA Sequencing: Prepare libraries with poly-A selection. Sequence on Illumina platform to a depth of 30-40 million 150bp paired-end reads per sample.
- Bioinformatic Analysis: Align reads to reference genome (GRCm38/GRCh38). Perform differential expression analysis (DESeq2, edgeR). Key Step: Compare differentially expressed genes (DEGs) from the targeted knockdown against the non-targeting control. Filter DEGs not predicted by TargetScan or off-target prediction algorithms (e.g., BLAST for seed region matches). Validate top off-target candidates by individual siRNA transfection.
Protocol: Monitoring Compensatory Pathway Activation via Phospho-Proteomics
Objective: To detect site-specific phosphorylation changes indicating activation of alternative signaling nodes following STAT3 knockdown in astrocytes.
Materials:
- Primary human astrocytes or U-251 MG cell line
- STAT3-specific CRISPR-Cas9 RNP complex (Alt-R CRISPR-Cas9 system)
- LC-MS/MS-grade reagents: urea, HEPES, protease/phosphatase inhibitors
- TMTpro 16plex kit for multiplexed quantitation
- TiO₂ or IMAC magnetic beads for phosphopeptide enrichment
- High-pH reverse-phase fractionation kit
Procedure:
- Genetic Perturbation: Transfect astrocytes with STAT3-targeting CRISPR RNP using electroporation (Neon System, 1400V, 20ms, 2 pulses). Include non-targeting gRNA control.
- Stimulation: At 96 hours post-editing (allow compensation to manifest), stimulate cells with IL-6 (50 ng/mL, 15 minutes) to activate JAK-STAT signaling.
- Cell Lysis and Digestion: Lyse cells in 8M urea buffer. Reduce with 5mM DTT, alkylate with 15mM iodoacetamide. Digest with Lys-C (1:100, 2h) followed by trypsin (1:50, overnight).
- TMT Labeling and Fractionation: Label digests with TMTpro tags according to kit instructions. Pool samples and fractionate using high-pH RP chromatography into 96 fractions consolidated into 24.
- Phosphopeptide Enrichment & LC-MS/MS: Enrich phosphopeptides from each fraction using TiO₂ beads. Analyze on Orbitrap Eclipse Tribrid MS with 3h gradient.
- Data Analysis: Process raw files with MaxQuant. Identify significantly altered phosphosites (p<0.01, fold-change >1.5) in the STAT3-KO vs control without IL-6 stimulation. Sites on proteins in related pathways (e.g., MAPK, NF-κB, other STATs) indicate compensation.
Protocol: Functional Rescue to Confirm Specificity
Objective: To distinguish phenotype causality from off-target/compensation effects by re-expressing an RNAi-resistant wild-type target gene.
Materials:
- HEK293T cells for virus production
- Target gene (e.g., SOCS1) cDNA clone in mammalian expression vector
- Site-directed mutagenesis kit to introduce silent mutations in siRNA target site
- Packaging plasmids (psPAX2, pMD2.G)
- Puromycin or other appropriate selection agent
Procedure:
- Construct RNAi-Resistant cDNA: Using the wild-type SOCS1 plasmid, perform site-directed mutagenesis to alter 4-6 nucleotides within the siRNA target region without changing the amino acid sequence (consult siRNA sequence from vendor).
- Produce Lentivirus: Co-transfect HEK293T cells with the resistant (or empty vector control) plasmid, psPAX2, and pMD2.G using PEI transfection reagent. Harvest virus-containing supernatant at 48 and 72 hours.
- Stable Cell Line Generation: Transduce target cells (e.g., microglia) with the virus, select with puromycin (2 µg/mL) for 7 days.
- Knockdown in Rescued Cells: Transfert the stable cell line (expressing RNAi-resistant SOCS1) with SOCS1-targeting siRNA or a non-targeting control.
- Assay: Measure downstream endpoints (e.g., STAT1 phosphorylation by Western blot, cytokine secretion by ELISA). Interpretation: If the phenotype from the original knockdown (e.g., hyperactive STAT1) is reversed in cells expressing the resistant cDNA but not the empty vector, the phenotype is likely on-target.
Visualization of Pathways and Workflows
Diagram Title: JAK-STAT Signaling and Compensatory Activation After Knockout
Diagram Title: Workflow for Addressing Off-Target and Compensation Effects
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Rigorous JAK-STAT Perturbation Studies
| Reagent Category | Specific Example(s) | Function & Rationale for Use |
|---|---|---|
| Validated Control Perturbations | Dharmacon ON-TARGETplus Non-targeting Control #1, Santa Cruz Biotechnology sc-37007 (shRNA), Addgene #105434 (non-targeting sgRNA) | Provides baseline for distinguishing true off-target effects from experimental noise; essential for RNA-seq validation. |
| Orthogonal Validation Tools | CRISPR-Cas9 KO for siRNA KD validation (and vice versa); dCas9-KRAB (CRISPRi) for transcriptional repression; dCas9-VPR (CRISPRa) for activation. | Confirms phenotype is specific to gene manipulation, not the method. CRISPRi/a avoids INDELs and DNA damage response. |
| Pathway-Specific Chemical Inhibitors | Ruxolitinib (JAK1/2), Tofacitinib (JAK1/3), Stattic (STAT3 SH2 domain), S3I-201 (STAT3 inhibitor). | Used in compensation assays to block suspected alternative pathways and test if they are sustaining the phenotype post-KO. |
| RNAi-Resistant cDNA Constructs | Custom gene synthesis or site-directed mutagenesis kits (e.g., NEB Q5) to create silent mutations in siRNA target site. | Gold-standard for rescue experiments to confirm on-target causality of observed phenotypes. |
| Multiplexed Readout Kits | Luminex xMAP cytokine/phosphoprotein panels, Proteome Profiler Phospho-Kinase Array (R&D Systems), TMTpro 16plex for proteomics. | Enables broad, simultaneous monitoring of pathway nodes and cytokines to detect compensatory shifts. |
| High-Fidelity Editing Systems | Alt-R S.p. HiFi Cas9 Nuclease V3 (IDT), TrueCut Cas9 Protein v2 (Thermo Fisher). | Reduces off-target editing compared to wild-type Cas9, crucial for clean knockout studies. |
| Bioinformatic Analysis Suites | CRISPOR (sgRNA design), DESeq2/edgeR (RNA-seq), MaxQuant (proteomics), PhosphoSitePlus (phosphosite analysis). | Essential for designing specific guides and rigorously analyzing omics data to identify off-targets/compensation. |
Best Practices for Reproducible Measurement of JAK-STAT Activity in Complex CNS Tissues
Within neuroinflammation research, the JAK-STAT signaling pathway is a critical mediator of glial cell activation, cytokine communication, and neuronal responses. Achieving reproducible quantification of its activity in the complex cellular milieu of the central nervous system (CNS) presents significant technical challenges. This guide outlines best practices for reliable measurement, framed within the mechanistic thesis of JAK-STAT activation as a driver of neuroinflammatory cascades.
Critical Challenges in CNS JAK-STAT Measurement
Key Obstacles:
- Cellular Heterogeneity: Signal from neurons, astrocytes, microglia, and oligodendrocytes must be distinguished.
- Low and Transient Phosphorylation: pSTAT is often low-abundance and temporally dynamic.
- Rapid Post-Mortem Changes: Phospho-epitope degradation during tissue harvest.
- Antibody Specificity: Cross-reactivity and validation in multiplex assays.
Quantitative Summary of Signal Dynamics:
| Parameter | Typical Range/Value | Notes for CNS Tissue |
|---|---|---|
| STAT1/3 Phosphorylation Peak | 5-30 min post-cytokine stimulus | Varies by cell type; microglia respond fastest. |
| Post-Mortem Delay Impact | Significant pSTAT loss after >5 min | Perfusion fixation is gold standard. |
| Effective Cytokine Dose (in vivo) | IL-6: 5-50 µg/kg; IFN-γ: 10-100 U/g | Dose-response is pathway-specific. |
| Signal-to-Noise in IHC | 2:1 to 10:1 (pSTAT:total STAT) | Highly dependent on fixation and retrieval. |
Experimental Protocols for Key Assays
Protocol: Perfusion Fixation for Optimal pSTAT Preservation
This protocol is critical for preserving the in vivo phosphorylation state.
- Anesthesia: Deeply anesthetize rodent with sodium pentobarbital (i.p., 50 mg/kg).
- Perfusion: Transcardially perfuse with 50 mL ice-cold 0.1 M PBS (pH 7.4), followed immediately by 100 mL of ice-cold 4% paraformaldehyde (PFA) in PBS.
- Brain Extraction & Post-Fixation: Dissect brain rapidly (<2 min) and place in same 4% PFA for 24 hours at 4°C.
- Cryoprotection: Transfer to 30% sucrose in PBS until tissue sinks (2-3 days).
- Sectioning: Snap-freeze in OCT compound. Cut 20-40 µm coronal sections on a cryostat.
Protocol: Multiplex Immunofluorescence for Cell-Type-Specific pSTAT
Enables co-localization of pSTAT with cell-specific markers.
- Section Prep: Wash free-floating or slide-mounted sections (from 2.1) in PBS.
- Antigen Retrieval: Use citrate buffer (pH 6.0) at 85°C for 20 min. Cool for 30 min.
- Blocking: Block in 10% normal donkey serum + 0.3% Triton X-100 in PBS for 2 hours.
- Primary Antibody Incubation: Incubate in cocktail for 48-72 hours at 4°C. Example: Chicken anti-GFAP (1:1000), Rabbit anti-pSTAT3 (Y705) (1:500), Mouse anti-NeuN (1:500).
- Secondary Incubation: Incubate in species-specific Alexa Fluor-conjugated antibodies (1:500) for 2 hours at RT.
- Imaging & Quantification: Acquire on a confocal microscope. Use automated image analysis software to quantify pSTAT intensity within defined cell masks (e.g., GFAP+ area).
Protocol: Phospho-Protein Immunoblot from CNS Sub-regions
For biochemical quantification from specific brain areas.
- Rapid Dissection: Following rapid decapitation without perfusion (for total protein), dissect hippocampus/cortex within 60 seconds and freeze in liquid N₂.
- Lysis: Homogenize in RIPA buffer supplemented with PhosSTOP phosphatase inhibitors and complete protease inhibitors. Centrifuge at 15,000g for 15 min at 4°C.
- Immunoblot: Resolve 20-40 µg protein on 8% SDS-PAGE. Transfer to PVDF. Blot sequentially: pSTAT1 (Y701), pSTAT3 (Y705), then total STAT1/STAT3, and finally β-Actin as a loading control. Use fluorescent secondary antibodies for linear quantitation.
Visualization of Core Concepts
Diagram 1: Core JAK-STAT Activation in Neuroinflammation.
Diagram 2: Experimental Workflow for CNS JAK-STAT Analysis.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Primary Function | Key Consideration for CNS |
|---|---|---|
| Phosphatase Inhibitors (e.g., PhosSTOP) | Preserve labile phospho-epitopes (pSTAT) during lysis. | Essential for biochemical assays; use at 2x recommended concentration. |
| Paraformaldehyde (4%, fresh) | Cross-linking fixative for immunohistochemistry. | Perfusion delivery is critical. pH must be 7.4. |
| Citrate Buffer (pH 6.0) | Heat-induced epitope retrieval solution for IHC. | Optimal for unmasking pSTAT epitopes; time/temp must be standardized. |
| Validated pSTAT Antibodies | Detect specific phosphorylated STATs (e.g., pSTAT1 Y701). | Must be validated for multiplex IHC. Lot-to-lot variation is common. |
| Cell-Type-Specific Markers | Identify neural cells (e.g., Iba1 for microglia, GFAP for astrocytes). | Required to assign JAK-STAT activity to specific CNS cell populations. |
| Fluorescent-Conjugated Secondaries | Enable multiplex detection and high-resolution imaging. | Use cross-adsorbed antibodies to prevent off-target labeling. |
| SOCS3 Reporter Constructs | Functional readout of JAK-STAT pathway activity via luciferase. | Useful in ex vivo slice cultures or primary glial cultures. |
| JAK-STAT Pathway Inhibitors (e.g., Ruxolitinib) | Pharmacological validation of signal specificity. | Determine CNS penetration; use in vivo controls for off-target effects. |
Validating the Target: Comparative Analysis and Clinical Translation of JAK-STAT Modulation
The Janus kinase-signal transducer and activator of transcription (JAK-STAT) pathway is a critical signaling cascade in neuroinflammation, with distinct activation patterns and functional outcomes in multiple sclerosis (MS), Alzheimer's disease (AD), and stroke. This review synthesizes current evidence, detailing pathway mechanisms, cytokine interactions, and cell-type-specific responses. Quantitative data are tabulated, and standard experimental protocols for pathway analysis in neuroinflammatory contexts are provided.
Within the broader thesis framework on JAK-STAT activation mechanisms in neuroinflammation, this analysis compares and contrasts the pathway's role across three major CNS disorders. The central thesis posits that while core JAK-STAT machinery is conserved, disease-specific cellular contexts, cytokine milieus, and temporal dynamics lead to divergent pro- or anti-inflammatory outcomes, dictating therapeutic targeting strategies.
Activation is triggered by extracellular cytokines (e.g., IFNs, IL-6, IL-10) binding to their cognate receptors, inducing receptor dimerization and trans-phosphorylation of associated JAKs. JAKs then phosphorylate receptor tails, creating docking sites for STAT monomers. Upon STAT phosphorylation, they dimerize, translocate to the nucleus, and regulate gene transcription. In the CNS, this occurs in microglia, astrocytes, oligodendrocytes, and neurons, influencing immune responses, glial activation, and neuronal survival.
Comparative Analysis: MS, Alzheimer's, and Stroke
Table 1: Key Cytokines and Primary JAK-STAT Components Involved
| Disease | Upstream Cytokines | Primary JAKs Activated | Primary STATs Activated | Predominant Cellular Source | Net Effect on Neuroinflammation |
|---|---|---|---|---|---|
| Multiple Sclerosis | IFN-γ, IL-12, IL-23 | JAK1, JAK2, TYK2 | STAT1, STAT3, STAT4 | Infiltrating T cells, Microglia | Pro-inflammatory (Th1/Th17 drive) |
| Alzheimer's Disease | IFN-γ, IL-4, IL-13, IL-10 | JAK1, JAK2, JAK3 | STAT1, STAT3, STAT6 | Microglia, Astrocytes | Dual: Pro- & Anti-inflammatory |
| Stroke (Ischemic) | IL-6, IFN-γ, IL-10 | JAK1, JAK2 | STAT1, STAT3, STAT5 | Microglia, Astrocytes, Neurons | Acute Pro-inflammatory, Later Repair |
Table 2: Key Transcriptional Targets and Functional Outcomes
| Disease | Key STAT Target Genes | Functional Consequence | Evidence from Preclinical Models |
|---|---|---|---|
| MS (EAE) | Nos2, Ccl5, Il12rb1, Mhc-II | Enhanced APC function, T cell infiltration, Demyelination | STAT1 KO→ Reduced EAE severity |
| Alzheimer's | Trem2, Arg1, Gfap, Bace1 | Altered phagocytosis, Aβ clearance, Astrogliosis | STAT3 inhibition→ Reduced gliosis, improved cognition |
| Stroke | Cox2, Mmp9, Vegfa, Bcl2 | BBB disruption, Inflammation, Angiogenesis, Cell survival | STAT3 inhibitor→ Smaller infarct size |
Multiple Sclerosis (MS)
In MS and its animal model (EAE), the JAK-STAT pathway is a primary driver of pathogenic T helper cell differentiation and CNS infiltration. IFN-γ/STAT1 and IL-12/STAT4 axes promote Th1 cells, while IL-23/STAT3 drives Th17 cells. Microglial STAT1 activation enhances antigen presentation and pro-inflammatory mediator release.
Experimental Protocol: Assessing STAT Phosphorylation in EAE CNS Tissue
- Induction: Induce EAE in C57BL/6 mice using MOG35-55 peptide emulsified in complete Freund's adjuvant, with pertussis toxin.
- Tissue Harvest: At peak disease (clinical score ~3-4), perfuse mice with ice-cold PBS. Dissect spinal cord and homogenize in RIPA buffer with phosphatase/protease inhibitors.
- Immunoblotting: Resolve 30 µg protein by SDS-PAGE, transfer to PVDF membrane. Block with 5% BSA. Incubate with primary antibodies: p-STAT1 (Tyr701), p-STAT3 (Tyr705), total STAT1, total STAT3, and β-actin loading control (all 1:1000, 4°C overnight). Use HRP-conjugated secondary antibodies (1:2000, 1h RT) and chemiluminescent detection.
- Analysis: Densitometry of bands normalized first to total STAT, then to β-actin. Compare to sham-immunized controls.
Alzheimer's Disease (AD)
In AD, JAK-STAT signaling exhibits complex, phase-dependent roles. In microglia, IFN-γ/STAT1 promotes a pro-inflammatory phenotype, potentially impairing Aβ clearance. Conversely, IL-4/STAT6 and IL-10/STAT3 can induce alternative activation, supporting clearance. Neuronal STAT3 may contribute to synaptic plasticity and survival.
Stroke (Ischemic)
Post-stroke, JAK-STAT has a dual temporal role. Early activation (hours-days) of STAT1 and STAT3 in microglia and astrocytes exacerbates inflammation and BBB breakdown. In the subacute phase (days-weeks), STAT3 in astrocytes and neurons promotes protective gliosis, angiogenesis, and neuronal survival, facilitating repair.
Experimental Protocol: Cell-Type Specific STAT3 Deletion in Stroke
- Mouse Models: Cross Stat3 floxed mice (Stat3^(fl/fl)) with cell-specific Cre drivers: Cx3cr1-CreER^T for microglia, Gfap-CreER^T for astrocytes, or CamkIIa-Cre for forebrain neurons. Induce Cre with tamoxifen (for ER^T models) prior to stroke.
- Stroke Surgery: Perform transient middle cerebral artery occlusion (tMCAO): insert a silicone-coated monofilament into the internal carotid artery to occlude the MCA for 60 minutes, then reperfuse.
- Assessment: At 24h (acute) and 7-28d (recovery) post-stroke, evaluate infarct volume (TTC staining), neurological deficit scores, and immunohistochemistry for cell markers (Iba1, GFAP, NeuN) and p-STAT3. Use flow cytometry on dissociated brain to analyze p-STAT3 in sorted cell populations.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Neuroinflammation Research
| Reagent | Example (Supplier) | Primary Function in Experiments |
|---|---|---|
| Phospho-specific STAT Antibodies | Anti-pSTAT1 (Tyr701) (Cell Signaling #7649) | Detecting pathway activation via WB, IHC, Flow |
| JAK-STAT Pathway Inhibitors | Tofacitinib (JAK1/3 inhibitor), Ruxolitinib (JAK1/2 inhibitor) (Selleckchem) | Pharmacological inhibition to establish causality |
| Cytokine Recombinant Proteins | Mouse IFN-γ, IL-6, IL-4 (PeproTech) | Stimulating pathway in vitro (glia/neuron cultures) |
| STAT Reporter Cell Lines | HEK-STAT1 or STAT3 Luciferase Reporter (BPS Bioscience) | High-throughput screening for pathway modulators |
| siRNA/shRNA Kits | Stat3 siRNA pools (Dharmacon) | Knockdown of specific components in cell culture |
| Multiplex Cytokine Assay | MILLIPLEX MAP Mouse Cytokine/Chemokine Panel (MilliporeSigma) | Profiling upstream cytokine milieu in tissue/biofluid |
| Chromatin Immunoprecipitation (ChIP) Kits | MAGnify ChIP Kit (Invitrogen) | Identifying direct STAT target gene promoters |
Discussion and Therapeutic Implications
The comparative analysis underscores that JAK-STAT inhibition is a promising strategy in MS (as evidenced by the success of oral JAK inhibitors) and the acute phase of stroke, but requires precise timing and cell-specific targeting in AD and the stroke recovery phase, where certain STAT functions are beneficial. Future research must leverage single-cell transcriptomics and spatial proteomics to resolve cell- and spatiotemporal-specific signaling networks for next-generation therapeutics.
Within the mechanistic thesis of JAK-STAT pathway activation in neuroinflammation, a critical pillar of validation is human genetic evidence. The pathway's role in cytokine signaling positions it centrally in neuroinflammatory and neuroimmune diseases. While in vitro and in vivo models establish mechanism, population-scale genomic studies provide causal inference, linking specific genetic variants in JAK-STAT genes to disease risk in humans. This guide details the methodologies, findings, and translational toolkit derived from these studies, validating the pathway as a therapeutic target.
Large-scale genome-wide association studies (GWAS) and whole-exome/genome sequencing (WES/WGS) have identified germline variants in JAK-STAT pathway genes associated with increased or decreased risk for immune-mediated and neuroinflammatory diseases. The table below summarizes recent, high-impact findings.
Table 1: Human Genetic Variants in JAK-STAT Pathway Genes Linked to Disease Risk
| Gene | Variant (rsID or Description) | Associated Phenotype | Study Type | Odds Ratio / Hazard Ratio (95% CI) | P-value | Proposed Functional Consequence |
|---|---|---|---|---|---|---|
| TYK2 | rs34536443 (P1104A) | Multiple Sclerosis (MS) | GWAS | 0.65 (0.59–0.71) | 2.0 × 10-18 | Loss-of-function; protective via reduced IL-12/IL-23 signaling. |
| JAK2 | rs77375493 (V625F) | Autoimmune Disorders (e.g., RA, SLE) | WES | 2.1 (1.7–2.6) | 4.3 × 10-12 | Gain-of-function; hyperactive signaling. |
| STAT4 | rs7574865 (intronic) | Rheumatoid Arthritis, SLE | GWAS | 1.32 (1.26–1.38) | 5.6 × 10-22 | Increased STAT4 expression/enhanced phosphorylation. |
| JAK1 | rs310241 (E322K) | Ulcerative Colitis | GWAS & Fine-mapping | 1.15 (1.11–1.19) | 8.9 × 10-14 | Modest gain-of-function in IL-6/IFN signaling. |
| STAT3 | Rare LoF variants | Autoimmune Disease (APDS-like) | WES/WGS | N/A (High Penetrance) | N/A | Loss-of-function; immune dysregulation. |
| SOCS1 | rs243327 (5'UTR) | Alzheimer's Disease (AD) | GWAS (immune subset) | 1.08 (1.05–1.11) | 3.1 × 10-8 | Reduced SOCS1 expression; disinhibited neuroinflammatory signaling. |
Detailed Experimental Protocols for Validation
Primary GWAS & Sequencing Analysis Workflow
Protocol Title: Case-Control GWAS for JAK-STAT Variant Discovery Objective: Identify common and rare genetic variants associated with disease risk.
- Cohort Ascertainment: Recruit large, well-phenotyped case and control cohorts with informed consent. Match for ancestry (e.g., via principal component analysis).
- Genotyping: Use high-density SNP arrays (e.g., Illumina Global Screening Array). Perform rigorous quality control (QC): call rate >98%, Hardy-Weinberg equilibrium P > 1×10-6, minor allele frequency (MAF) checks.
- Imputation: Impute to a reference panel (e.g., 1000 Genomes, TOPMed) to increase variant coverage. Retain well-imputed variants (INFO score >0.8).
- Association Testing: Perform logistic regression for each variant, adjusting for principal components, sex, and other relevant covariates. Apply genome-wide significance threshold (P < 5×10-8).
- Replication: Test top-associated signals in an independent replication cohort.
- Fine-mapping: For significant loci, use statistical methods (e.g., SuSiE) to identify likely causal variants from a set of correlated SNPs.
In VitroFunctional Validation of Identified Variants
Protocol Title: Luciferase Reporter Assay for Enhancer/ Promoter Variants Objective: Determine if a non-coding GWAS variant alters transcriptional activity.
- Cloning: Amplify genomic DNA regions (~500-1000bp) containing the reference and alternative alleles of the SNP from human genomic DNA. Clone into a luciferase reporter vector (e.g., pGL4.23) upstream of a minimal promoter.
- Cell Culture & Transfection: Culture relevant cell lines (e.g., HEK293T, Jurkat T-cells, human microglia cell line HMC3). Seed cells in 24-well plates. Co-transfect reporter construct and a Renilla luciferase control plasmid (e.g., pRL-TK) for normalization using a suitable transfection reagent.
- Stimulation: If relevant, stimulate cells with cytokines (e.g., IFN-γ, IL-6) known to act through the JAK-STAT pathway 24h post-transfection.
- Luciferase Assay: Harvest cells 48h post-transfection. Measure firefly and Renilla luciferase activity using a dual-luciferase assay kit. Normalize firefly luminescence to Renilla.
- Analysis: Perform experiment in triplicate, repeated at least three times. Compare normalized luminescence between alleles using a Student's t-test.
Visualizing the Genetic Evidence within the Neuroinflammatory Pathway
Diagram 1: JAK-STAT Pathway in Neuroinflammation & Key Genetic Hits
Diagram 2: Experimental Validation Workflow for GWAS Hits
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Validating JAK-STAT Genetic Variants
| Reagent Category | Specific Example | Function & Application |
|---|---|---|
| Genotyping & Sequencing | Illumina Infinium Global Screening Array-24 v3.0 | High-throughput genotyping of common SNPs for GWAS cohort building. |
| IDT xGen Hybridization Capture Probes (JAK-STAT gene panel) | Targeted enrichment for deep sequencing of JAK-STAT genes in cases/controls. | |
| Cell-Based Assays | Phospho-STAT3 (Tyr705) Alexa Fluor 488 Conjugate Antibody (CST) | Flow cytometry to measure STAT activation kinetics in primary immune cells from variant carriers. |
| Dual-Luciferase Reporter Assay System (Promega) | Quantifying allele-specific effects of non-coding variants on promoter/enhancer activity. | |
| Recombinant Human Cytokines (IFN-γ, IL-6, IL-23) (R&D Systems) | Stimulation of the JAK-STAT pathway in functional assays. | |
| Genetic Engineering | CRISPR-Cas9 Gene Editing System (Synthego) | Isogenic cell line generation (knock-in of risk/protective variant) for clean functional comparison. |
| Allele-Specific qPCR Probes (TaqMan) | Quantifying allele-specific expression (ASE) in heterozygous samples to assess regulatory impact. | |
| Pathway Modulation | JAK Inhibitors (e.g., Tofacitinib, Ruxolitinib) (Selleckchem) | Pharmacological tools to probe pathway necessity and rescue phenotypes in hyperactive variants. |
| Primary Tissue Models | Cryopreserved Human PBMCs from Genotyped Donors (STEMCELL) | Ex vivo analysis of signaling and cytokine production linked directly to donor genotype. |
Within the thesis on JAK-STAT pathway mechanism in neuroinflammation, analyzing the transition of Janus kinase inhibitors (JAKi) from preclinical models to clinical trials is paramount. Neuroinflammatory disorders, such as multiple sclerosis (MS), Alzheimer's disease, and stroke, involve dysregulated activation of the JAK-STAT signaling cascade in glial cells and neurons. This whitepaper provides a technical guide for researchers on critically evaluating the concordance or disconnect between efficacy and safety signals of JAKi across experimental and human studies.
The JAK-STAT Pathway in Neuroinflammation: A Mechanistic Primer
Activation begins with extracellular cytokine binding (e.g., IL-6, IFN-γ) to its receptor, inducing JAK auto-phosphorylation and subsequent phosphorylation of STAT proteins. Phosphorylated STATs dimerize, translocate to the nucleus, and drive transcription of pro-inflammatory genes. In neurological contexts, microglial and astrocytic JAK-STAT hyperactivation perpetuates inflammation and neuronal damage.
Diagram Title: JAK-STAT Activation in Neuroinflammation
Preclinical Data: Efficacy and Safety Models
3.1 Key Experimental Models:
- In Vitro: Primary rodent/human microglia or astrocyte cultures stimulated with cytokines (IFN-γ, IL-6).
- In Vivo: Experimental Autoimmune Encephalomyelitis (EAE) for MS, LPS-induced neuroinflammation, ischemic stroke models (MCAO), and transgenic Alzheimer's models (APP/PS1).
3.2 Typical Efficacy Endpoints:
- Reduction in phosphorylated STAT levels in CNS tissue (Western blot, IHC).
- Attenuation of pro-inflammatory gene expression (IL-1β, TNF-α, iNOS) via qPCR.
- Improved clinical score (e.g., EAE limb paralysis).
- Histopathological improvement: reduced demyelination, microgliosis, neuronal loss.
3.3 Safety & Pharmacokinetic Assessments:
- Plasma and brain concentration (Kp,uu) via LC-MS/MS.
- Hematological profiles (CBC), liver enzymes (ALT/AST).
- Off-target kinase screening panels.
Table 1: Summary of Preclinical JAKi Findings in Neurological Models
| JAKi (Example) | Model (Species) | Primary Efficacy Outcome | Dose/Route | Key Safety Finding | Brain Penetrance (Kp,uu) |
|---|---|---|---|---|---|
| Tofacitinib (Pan-JAKi) | EAE (Mouse) | 60% reduction in clinical score, 70% ↓ pSTAT3 in spinal cord | 30 mg/kg, oral, BID | Mild lymphopenia | 0.1-0.3 |
| Ruxolitinib (JAK1/2i) | LPS Neuroinflammation (Rat) | 80% ↓ TNF-α mRNA in cortex | 10 mg/kg, i.p., QD | Transient anemia | 0.05 |
| Upadacitinib (JAK1i) | MCAO (Mouse) | 40% reduction in infarct volume | 5 mg/kg, oral, QD | No significant change in platelets | 0.15 |
| Experimental JAKi-X | APP/PS1 (Mouse) | 50% ↓ Aβ plaque-associated microgliosis | 15 mg/kg, oral, BID | Elevated liver enzymes at high dose | 0.8 |
3.4 Detailed Protocol: Assessing pSTAT in EAE Spinal Cord
- Animal Model: C57BL/6 mice immunized with MOG35-55/CFA.
- Treatment: JAKi or vehicle from day 7 post-immunization.
- Tissue Collection: At clinical peak, perfuse with PBS. Harvest spinal cord, snap-freeze.
- Western Blot:
- Homogenize tissue in RIPA buffer with protease/phosphatase inhibitors.
- Resolve 30 µg protein on 4-12% Bis-Tris gel.
- Transfer to PVDF membrane.
- Block with 5% BSA/TBST for 1h.
- Incubate with primary antibodies: anti-pSTAT3 (Tyr705) (1:1000) and anti-STAT3 (1:2000) overnight at 4°C.
- Incubate with HRP-conjugated secondary antibody (1:5000) for 1h.
- Develop with ECL substrate, image, and quantify band density relative to total STAT3.
Clinical Trial Data: Translation to Humans
4.1 Completed & Ongoing Trials: Examples include tofacitinib in MS (NCT04035005), ruxolitinib in Alzheimer's (NCT04673162).
4.2 Efficacy Endpoints:
- Phase II: Biomarker changes (CSF pSTAT levels, neurofilament light chain), MRI lesion activity (MS), cognitive scales (ADAS-Cog in AD).
- Phase III: Clinical disability progression (EDSS in MS), composite cognitive/functional scores.
4.3 Safety Monitoring:
- Routine: Infections, hematology, hepatic panels.
- JAKi-class specific: Major adverse cardiac events (MACE), thromboembolism (VTE), malignancy risk.
Table 2: Clinical Trial Data Snapshot for JAKi in Neurological Indications
| JAKi | Trial Phase | Indication | Primary Efficacy Result (vs. Placebo) | Notable Safety Signals | Reference (Example) |
|---|---|---|---|---|---|
| Tofacitinib | Phase 2 | Relapsing MS | 45% reduction in new MRI lesions at 24 weeks | Increased herpes zoster infections, minor LDL increase | Lancet Neurol 2021 |
| Ruxolitinib | Phase 2 | Alzheimer's | No significant change in ADAS-Cog at 24 weeks | Anemia (dose-dependent), no MACE imbalance | Published Abstract |
| JAKi-Y | Phase 3 | Ischemic Stroke | Ongoing (P: Change in mRS at 90 days) | Monitoring for infections & VTE | NCTXXXXXXX |
Critical Analysis: Preclinical-Clinical Disconnect
5.1 Efficacy Gaps:
- Species-specific differences in JAK-STAT biology and disease pathology.
- Temporal Factor: Acute preclinical models vs. chronic human disease.
- Biomarker Translation: Reduction in CNS pSTAT in rodents may not correlate with human CSF pSTAT or clinical benefit.
5.2 Safety Gaps:
- Preclinical toxicology often misses immune-mediated risks (e.g., specific infection susceptibilities).
- Cardiovascular and thromboembolic risks emerge only in large, long-term human trials.
Diagram Title: Preclinical vs Clinical Data Translation Gaps
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Neuroinflammation Research
| Item | Function/Brief Explanation | Example Vendor/Cat. No.* |
|---|---|---|
| Phospho-STAT Specific Antibodies | Detect activated STATs via WB/IHC/Flow. Critical for measuring pathway inhibition. | Cell Signaling Technology #9145 (pSTAT3 Tyr705) |
| Selective JAK Inhibitors (Tool Compounds) | For in vitro and in vivo mechanistic studies. Validate target engagement. | MedChemExpress (HY-40354 for Tofacitinib) |
| Cytokine Stimulation Kits | Standardized in vitro activation of JAK-STAT in glial/neuronal cultures. | R&D Systems, Human/Mouse IFN-γ |
| Multiplex Immunoassay Panels | Quantify panels of cytokines/chemokines in cell supernatant, serum, or brain homogenate. | Meso Scale Discovery (Proinflammatory Panel 1) |
| Kinase Profiling Assay Services | Assess selectivity of novel JAKi against large kinase panels to predict off-target effects. | Eurofins DiscoverX KinomeScan |
| Brain Homogenization Kits | With protease/phosphatase inhibitors for optimal phospho-protein preservation. | Thermo Fisher #87786 |
| Barrier-Specific Cell Lines | (e.g., hCMEC/D3) to model blood-brain barrier permeability of JAKi in vitro. | Merck #SCC066 |
Examples for illustration; not endorsements.
This whitepaper is framed within a broader thesis investigating the spatiotemporal dysregulation of the JAK-STAT pathway as a core mechanism of activation in neuroinflammatory cascades. In the CNS, resident microglia, astrocytes, and infiltrating immune cells utilize distinct JAK-STAT isoform combinations for cytokine signaling, making the pharmacological selectivity profile of JAK inhibitors (JAKi) a critical determinant of therapeutic efficacy and safety in neurological diseases.
JAK-STAT Isoforms and CNS Expression Profiles
The four JAK isoforms (JAK1, JAK2, JAK3, TYK2) and seven STAT isoforms exhibit cell-type-specific expression and function in the healthy and inflamed CNS.
Table 1: Primary JAK-STAT Isoform Pairings for Neuroinflammatory Cytokines
| Cytokine/Signal | Primary Receptor Complex | JAK Isoforms Engaged | STAT Isoform Activated | Primary CNS Cellular Target |
|---|---|---|---|---|
| IFN-γ | IFNGR1/IFNGR2 | JAK1, JAK2 | STAT1, STAT3, STAT5 | Microglia, Astrocytes |
| IL-6 | IL-6Rα/gp130 | JAK1, JAK2, TYK2 | STAT3, STAT1 | Astrocytes, Neurons |
| GM-CSF | GM-CSFRα/βc | JAK2, TYK2 | STAT5, STAT3 | Microglia |
| IL-4/IL-13 | Type II IL-4R | JAK1, JAK3, TYK2 | STAT6 | Microglia (M2 Polarization) |
| IFN-α/β | IFNAR1/IFNAR2 | JAK1, TYK2 | STAT1, STAT2, STAT3 | All Neural Cells |
Pharmacological Selectivity Profiles: Quantitative Data
Inhibitors are classified by their selectivity for JAK isoforms, quantified by IC50 or in vitro kinase assay data.
Table 2: Selectivity Profiles of Pan-JAK vs. Isoform-Selective Inhibitors (Representative Compounds)
| Inhibitor Class | Example Compound(s) | JAK1 IC50 (nM) | JAK2 IC50 (nM) | JAK3 IC50 (nM) | TYK2 IC50 (nM) | Key CNS Application (Research/Clinical) |
|---|---|---|---|---|---|---|
| Pan-JAK | Tofacitinib, Peficitinib | 112 (JAK1) | 20 (JAK2) | 1.4 (JAK3) | 34 (TYK2)* | Broad-spectrum neuroinflammation models (e.g., EAE) |
| JAK1-Selective | Upadacitinib, Filgotinib | 43 (JAK1) | 200 (JAK2) | 1250 (JAK3) | 1800 (TYK2) | Targeting IL-6 & IFN-γ signaling in astrocytes |
| JAK2-Selective | Fedratinib, BMS-911543 | 829 (JAK1) | 3 (JAK2) | 330 (JAK3) | 668 (TYK2) | Microglial activation models (GM-CSF dependent) |
| TYK2-Selective | Deucravacitinib, BMS-986165 | >10,000 (JAK1) | >10,000 (JAK2) | >10,000 (JAK3) | 0.2 (TYK2) | IFN-α/β driven pathologies (e.g., SLE neuropsychiatric) |
| JAK3-Selective | Decernotinib (VX-509) | 383 (JAK1) | 1300 (JAK2) | 29 (JAK3) | 2100 (TYK2) | Limited in CNS; used in peripheral immune cell studies |
Note: Tofacitinib's TYK2 inhibition is functional, though direct binding is weaker. IC50 values are approximate and assay-dependent.
Table 3: CNS Penetration Metrics (Rodent PK Studies)
| Compound | Brain-to-Plasma Ratio (Kp) | P-gp Substrate (Yes/No) | Primary Evidence of CNS Target Engagement |
|---|---|---|---|
| Tofacitinib | 0.1 - 0.3 | Yes | Reduction in pSTAT3 in hippocampal microglia after systemic LPS |
| Upadacitinib | 0.2 - 0.5 | Yes | Dose-dependent inhibition of IFN-γ-induced pSTAT1 in cortex |
| Fedratinib | ~1.0 | Weak | Inhibition of hippocampal neural progenitor cell pSTAT5 |
| Deucravacitinib | 0.15 | Yes | Allosteric inhibitor; high in vivo potency despite low Kp |
| AZD1480 (JAK1/2) | 0.8 - 1.2 | No | Robust inhibition of JAK/STAT in brain tumor microenvironment |
Core Signaling Pathways: Mechanism of Activation
The canonical JAK-STAT pathway activation in CNS cell types.
Diagram Title: Canonical JAK-STAT Activation and Inhibition in CNS
Experimental Protocols for Assessing Inhibitor Profiles in CNS Research
Protocol:In VitroSelectivity Profiling (Cell-Free Kinase Assay)
Objective: Determine IC50 values against purified human JAK isoforms.
- Reagents: Recombinant human JAK1, JAK2, JAK3, TYK2 kinase domains (SignalChem); ATP (1 mM stock); Substrate peptide (e.g., Poly(Glu4,Tyr1)); Detection reagent (ADP-Glo Kinase Assay, Promega).
- Procedure:
- Prepare inhibitor in 10-dose, 3-fold serial dilution in DMSO.
- In white 384-well plate, add 2.5 μL of kinase (2 nM final) in assay buffer.
- Add 0.5 μL of inhibitor dilution. Include DMSO-only controls (100% activity) and no-kinase controls (0% activity).
- Start reaction by adding 2 μL of ATP/peptide mixture (5 μM ATP, 0.2 mg/mL peptide final).
- Incubate at 25°C for 60 min.
- Terminate reaction with 5 μL ADP-Glo Reagent, incubate 40 min.
- Add 10 μL Kinase Detection Reagent, incubate 30 min, read luminescence.
- Analysis: Fit dose-response curves (log[inhibitor] vs. normalized response) using 4-parameter logistic model (e.g., GraphPad Prism) to calculate IC50.
Protocol: Cellular Target Engagement in Primary Murine Microglia
Objective: Measure inhibition of cytokine-specific STAT phosphorylation.
- Reagents: Primary microglia from C57BL/6J P1-3 pups; DMEM/F-12 complete medium; recombinant murine IFN-γ & GM-CSF (PeproTech); inhibitors solubilized in DMSO; Phospho-STAT1 (Tyr701) & Phospho-STAT5 (Tyr694) antibodies (Cell Signaling Technology); Flow cytometry buffer.
- Procedure:
- Seed 2x10^5 cells/well in 96-well plate. Pre-treat with inhibitor (0.1 nM - 10 μM) or vehicle (0.1% DMSO) for 60 min.
- Stimulate with IFN-γ (50 ng/mL) or GM-CSF (20 ng/mL) for 15 min.
- Aspirate medium, dissociate cells with gentle enzymatic treatment.
- Fix immediately with pre-warmed 4% PFA for 10 min at 37°C.
- Permeabilize with ice-cold 90% methanol, store at -20°C overnight.
- Stain with phospho-specific antibody (1:100) in flow buffer for 1 hr at RT.
- Analyze via flow cytometry. Gate on live single cells, measure geometric MFI of phospho-STAT signal.
- Analysis: Calculate % inhibition relative to vehicle-treated, cytokine-stimulated controls. Generate IC50 values for cellular potency.
Protocol:In VivoBrain Target Engagement (Rodent)
Objective: Assess brain penetration and inhibition of JAK-STAT pathway after systemic dosing.
- Reagents: Adult C57BL/6J mice; inhibitor formulated in 0.5% methylcellulose/0.1% Tween80; LPS (E. coli 055:B5); perfusion apparatus.
- Procedure:
- Administer inhibitor or vehicle via oral gavage (n=6/group). Use a pharmacokinetic-driven dosing regimen (e.g., 1 hr pre-challenge for peak plasma time).
- Administer LPS (1 mg/kg, i.p.) to induce systemic and CNS inflammation.
- Euthanize at predetermined time (e.g., 2h post-LPS). Collect blood via cardiac puncture for plasma. Transcardially perfuse with ice-cold PBS.
- Rapidly dissect brain regions (cortex, hippocampus). Hemisect: one half snap-frozen in liquid N2 for compound quantification (LC-MS/MS); the other half homogenized in RIPA buffer for phospho-STAT Western blot or Meso Scale Discovery (MSD) phospho-STAT multiplex assay.
- Analysis:
- PK: Calculate brain/plasma ratio (Kp). Determine brain unbound fraction (fu,brain) using equilibrium dialysis.
- PD: Quantify pSTAT levels. Relate brain unbound inhibitor concentration to % pathway inhibition for in vivo potency (EC50).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for JAK-STAT CNS Pharmacology Research
| Item | Function & Application | Example Product/Source |
|---|---|---|
| Selective JAK Inhibitors (Tool Compounds) | In vitro and in vivo pharmacological probes to dissect isoform-specific functions. | HY-40354 (Tofacitinib, MedChemExpress); S2789 (Upadacitinib, Selleckchem); BMS-911543 (Tocris). |
| Phospho-Specific STAT Antibodies | Detect pathway activation via WB, IHC, or flow cytometry. Critical for PD readouts. | Phospho-STAT3 (Tyr705) #9145; Phospho-STAT1 (Tyr701) #9167 (Cell Signaling Technology). |
| Multiplex Phospho-STAT Assay | Simultaneously quantify multiple pSTATs from limited tissue lysates (e.g., brain homogenates). | V-PLEX Phospho-STAT Panel 1 (STAT1/3/5) (Meso Scale Discovery). |
| Recombinant JAK Kinase Domains | For biochemical IC50 determination and screening. | Recombinant Human JAK1 (amino acids 866-1154) (SignalChem, #J01-10G). |
| Primary CNS Cell Kits | Isolate specific cell types for cell-autonomous signaling studies. | Primary Microglia Isolation Kit (Miltenyi Biotec, #130-110-634); Primary Astrocyte Media (ScienCell, #1801). |
| JAK-STAT Reporter Cell Lines | Stable cell lines for high-throughput screening of inhibitor activity in a cellular context. | HEK293 STAT1 or STAT3 Luciferase Reporter Cell Line (BPS Bioscience, #60610). |
| Brain Tissue Homogenization Kits | Efficient lysis for preserving labile phospho-epitopes from brain tissue. | Minute Total Protein Extraction Kit for Animal Tissues (Invent Biotechnologies, #AT-022). |
Comparative Efficacy and Safety Considerations in CNS
Table 5: Functional Outcomes in Preclinical Neuroinflammatory Models
| Inhibitor Class | EAE Model Efficacy (Max Clinical Score Reduction) | Impact on Microglial Phagocytosis | Impact on Oligodendrocyte Precursor Cell (OPC) Differentiation | Notable CNS Toxicity Concerns (Preclinical) |
|---|---|---|---|---|
| Pan-JAK (Tofacitinib) | 50-70% | Suppresses | Enhanced (via reduced inflammatory inhibition) | Lymphopenia, increased CNS viral load (e.g., JCV) |
| JAK1-Selective (Upadacitinib) | 40-60% | Mildly Suppresses | Mildly Enhanced | Anemia (mild), potential hepatotoxicity |
| JAK2-Selective (Fedratinib) | 20-40% | Potently Suppresses | Inhibited (via STAT5 blockade) | Cerebellar degeneration (thiamine-related), severe anemia |
| TYK2-Selective (Deucravacitinib) | 60-80% | Minimal effect | No direct effect | Favorable; minimal hematologic toxicity |
Diagram Title: Pan vs. Selective JAKi: Efficacy-Toxicity Trade-off Logic
The choice between pan-JAK and isoform-selective inhibitors in CNS disorders must be guided by the specific neuroinflammatory pathophysiology. Pan-JAK inhibitors offer broad suppression but carry a higher risk of disrupting homeostatic JAK-STAT functions critical for neural health. Selective agents, particularly JAK1 and TYK2 inhibitors, present a more refined tool, potentially improving the therapeutic window. Future research must prioritize the development of CNS-penetrant, isoform-selective inhibitors with optimized pharmacokinetic profiles and the validation of cell-specific biomarkers of target engagement to translate these pharmacological principles into effective neurotherapeutics.
Within neuroinflammation research, the JAK-STAT pathway is a critical signaling hub activated by cytokines like IL-6, IFN-γ, and others. Its dysregulation is implicated in multiple sclerosis, Alzheimer's disease, and other neuroinflammatory conditions. The phosphorylation of STAT (p-STAT) proteins and the subsequent feedback expression of Suppressors of Cytokine Signaling (SOCS) proteins are pivotal regulatory events. This technical guide provides a framework for validating these molecular events as robust, quantitative biomarkers for assessing treatment response to JAK-STAT-targeted therapies.
Core Biology: The JAK-STAT-SOCS Axis in Neuroinflammation
The canonical pathway is initiated when a cytokine binds its receptor, inducing JAK autophosphorylation and activation. Activated JAKs phosphorylate receptor tails, creating docking sites for STAT monomers. STATs are then phosphorylated, dimerize, and translocate to the nucleus to drive gene transcription, including that of SOCS genes. SOCS proteins then complete a negative feedback loop by inhibiting JAK kinase activity or targeting proteins for degradation.
Diagram: JAK-STAT-SOCS Signaling and Inhibition in Neuroinflammation
Table 1: Comparison of p-STAT and SOCS as Potential Biomarkers
| Parameter | p-STAT (e.g., p-STAT1, p-STAT3) | SOCS (e.g., SOCS1, SOCS3) |
|---|---|---|
| Molecular Nature | Post-translational modification (phosphorylation). | Induced protein expression (transcriptional feedback). |
| Kinetics | Rapid (minutes to 1-2 hours post-cytokine stimulation). Transient. | Delayed (1-4 hours post-stimulation). More sustained. |
| Stability in Samples | Labile; requires rapid fixation/phosphate inhibitors. | More stable; less sensitive to pre-analytical delay. |
| Detection Primary Methods | Phospho-flow cytometry, Western blot, IHC/IF with phospho-specific antibodies. | qRT-PCR, Western blot, RNA-Seq, IHC/IF. |
| Correlation with Activity | Direct measure of pathway activation flux. | Indirect measure of pathway activity via feedback strength. |
| Key Challenge | Pre-analytical variability; contextual (cell-type specific). | Specificity (regulated by other pathways); baseline variability. |
| Therapeutic Correlation | Direct: Reduction indicates successful JAK/STAT inhibition. | Indirect: Modulation may indicate effective pathway engagement or compensatory feedback. |
Detailed Experimental Protocols
Protocol: Phospho-STAT Analysis by Intracellular Flow Cytometry (Phospho-Flow)
Objective: Quantify cell-type-specific p-STAT levels in mixed CNS-derived immune cells (e.g., from murine EAE model or human PBMCs/CSF).
Materials & Reagents:
- Stimulus: Recombinant cytokine (e.g., IFN-γ, IL-6).
- Fixation: Pre-warmed 1.5% formaldehyde (UltraPure) or BD Cytofix.
- Permeabilization: 100% ice-cold methanol or commercial perm buffers (e.g., BD Phosflow Perm III).
- Antibodies: Conjugated anti-p-STAT1 (Y701), anti-p-STAT3 (Y705), anti-CD45, anti-CD11b, anti-CD3 for surface; viability dye.
- Controls: Unstimulated cells, cells + cytokine, cells + cytokine + JAK inhibitor (e.g., Tofacitinib).
Procedure:
- Single-cell suspension: Prepare cells in complete media. Rest for 1h at 37°C.
- Stimulation: Aliquot cells. Add cytokine (e.g., 50ng/mL IFN-γ) for 15 minutes. Include inhibitor control (pre-incubate 30min).
- Fixation: Immediately add equal volume pre-warmed fixative. Incubate 10min at 37°C.
- Permeabilization: Pellet, wash with PBS. Resuspend in ice-cold methanol. Incubate ≥30min at -20°C.
- Staining: Wash twice with staining buffer. Stain with surface antibodies (30min, RT, dark). Wash. Stain with intracellular p-STAT antibodies (60min, RT, dark).
- Acquisition: Acquire on a flow cytometer within 24h. Use unstimulated and fluorescence-minus-one (FMO) controls for gating.
Diagram: Phospho-STAT Flow Cytometry Workflow
Protocol: SOCS Expression Analysis by Quantitative RT-PCR
Objective: Measure SOCS1/SOCS3 mRNA induction as a functional readout of prior JAK-STAT activation in tissue or sorted cells.
Materials & Reagents:
- RNA Isolation: TRIzol or column-based kits (RNase-free).
- cDNA Synthesis: High-capacity cDNA reverse transcription kit with random hexamers.
- qPCR: SYBR Green or TaqMan master mix, primer/probe sets for SOCS1, SOCS3, and housekeeping genes (Gapdh, Hprt, Actb).
- Equipment: Thermal cycler, real-time PCR system.
Procedure:
- Sample Lysis: Homogenize tissue or lyse cell pellets in TRIzol. Store at -80°C or proceed.
- RNA Isolation: Follow manufacturer's protocol. Include DNase treatment. Quantify via Nanodrop.
- cDNA Synthesis: Use 500ng-1μg total RNA in 20μL reaction. Conditions: 25°C (10min), 37°C (120min), 85°C (5min).
- qPCR Setup: Prepare master mix with primers (e.g., 400nM each) and cDNA template (1:10 dilution). Run in triplicate.
- Cycling: Standard 2-step cycling (95°C denaturation, 60°C annealing/extension for 40 cycles).
- Analysis: Calculate ΔΔCt relative to housekeeping gene and untreated control.
Research Reagent Solutions & Essential Materials
Table 2: Scientist's Toolkit for JAK-STAT Biomarker Validation
| Reagent / Material | Function & Critical Notes |
|---|---|
| Phospho-Specific Antibodies (p-STAT1/3/5) | Detect active STATs. Must be validated for application (WB, flow, IHC). High lot-to-lot consistency is crucial. |
| SOCS1/SOCS3 Antibodies (for WB/IHC) | Detect feedback proteins. Often challenging for IHC; rigorous positive/negative controls required. |
| JAK Inhibitors (e.g., Tofacitinib, Ruxolitinib) | Pharmacological tool controls to confirm pathway-specific changes in p-STAT/SOCS. Use at published IC50 concentrations. |
| Recombinant Cytokines (IFN-γ, IL-6, OSM) | Pathway stimulants for ex vivo biomarker induction assays. Use carrier-free, high-purity grades. |
| Phosphatase & Protease Inhibitor Cocktails | Essential pre-analytical additives for tissue lysis buffers to preserve p-STAT signals. Must be fresh. |
| Viability Dye (e.g., Fixable Viability Stain) | Critical for flow cytometry to exclude dead cells which exhibit high non-specific phospho-signaling. |
| Single-Cell Isolation Kits (for CNS tissue) | Gentle enzymatic/mechanical dissociation kits to obtain viable single cells from brain/spinal cord for phospho-flow. |
| RNA Stabilization Reagent (e.g., RNAlater) | For SOCS mRNA analysis from tissues; inactivates RNases immediately upon collection. |
| TaqMan Gene Expression Assays | Pre-validated primer/probe sets for human/murine SOCS1, SOCS3. Provide superior specificity vs. SYBR Green for homologous genes. |
| Multiplex Luminex/Cytometry Bead Array | Platform for measuring multiple phospho-proteins or cytokines simultaneously from limited sample volumes (e.g., CSF). |
Validation Strategy & Data Interpretation
Validation requires a multi-tier approach:
- Analytical Validation: Establish assay precision, accuracy, sensitivity, linear range, and reproducibility for p-STAT and SOCS measurements.
- Biological Validation: Demonstrate correlation between biomarker level and in vivo pathway activity (using genetic or pharmacological modulators).
- Clinical/Biological Correlation: In pre-clinical models, correlate biomarker modulation with therapeutic efficacy (e.g., clinical score in EAE, histopathology). In human studies, correlate with treatment dose and clinical response.
Diagram: Biomarker Validation Logic Flow
Integrating p-STAT and SOCS measurements provides a complementary, dynamic readout of JAK-STAT pathway activity in neuroinflammation. p-STAT offers a direct, proximal snapshot of activation, while SOCS reflects integrated feedback. Robust validation of these biomarkers requires strict protocol standardization, careful reagent selection, and correlation with functional outcomes. When implemented rigorously, they can significantly enhance the evaluation of treatment response in both preclinical research and clinical trials for JAK-STAT-targeted neurotherapeutics.
The JAK-STAT pathway is a principal signaling cascade transducing extracellular cytokine signals into transcriptional programs within the central nervous system (CNS). In neuroinflammatory diseases—such as multiple sclerosis, Alzheimer's disease, and neuropathic pain—chronic activation of this pathway, particularly involving STAT1, STAT3, and STAT5, drives pathogenic processes including glial activation, immune cell infiltration, and neuronal apoptosis. While first-generation pan-JAK inhibitors have shown promise, their systemic immunosuppressive effects and lack of CNS penetrance limit therapeutic utility. This whitepaper details emerging strategies that target downstream STAT proteins with high selectivity and explores rational combination therapies to overcome pathway redundancy and resistance, thereby advancing a core thesis in neuroinflammation research: precise inhibition of specific STAT dimer species can modulate discrete neuroinflammatory gene networks with improved safety and efficacy.
Next-Generation Selective STAT Inhibitors: Mechanisms & Candidates
Next-generation inhibitors aim to disrupt STAT function through mechanisms beyond the traditional SH2 domain phosphotyrosine binding blockade. These include:
- Allosteric Inhibitors: Binding outside the SH2 domain to stabilize inactive conformations.
- DNA-Binding Disruptors: Preventing STAT dimer binding to gamma-activated sequence (GAS) elements.
- Dimerization Disruptors: Targeting the coiled-coil or N-domain interfaces.
- Transcriptional Complex Disruptors: Inhibiting interactions with co-activators like p300/CBP.
Table 1: Profiles of Next-Generation Selective STAT Inhibitors in Development
| STAT Target | Compound Name (Code) | Mechanism of Action | Development Stage (as of 2024) | Key Evidence in Neuroinflammation Models |
|---|---|---|---|---|
| STAT3 | BP-1-102 | SH2 domain allosteric binder, disrupts dimerization | Preclinical | Reduces astrogliosis and improves recovery in murine experimental autoimmune encephalomyelitis (EAE). |
| STAT3 | TTI-101 (C188-9) | Binds to STAT3 dimer interface, promotes degradation | Phase I/II Trials (oncology) | Shown to cross BBB*; suppresses microglia-mediated neurotoxicity in vitro. |
| STAT1 | Fludarabine | Inhibits STAT1 phosphorylation & DNA binding | Approved (oncology), Repurposing | Attenuates IFN-γ-induced microglial activation and iNOS production. |
| STAT5 | AC-4-130 | Disrupts STAT5 dimerization via SH2 domain binding | Preclinical | Limits oligodendrocyte precursor cell apoptosis induced by pro-inflammatory cytokines. |
| STAT3/STAT1 | Napabucasin | Inhibits STAT3-driven gene transcription (p-STAT3 shuttling) | Phase III Trials (oncology) | Demonstrated efficacy in glioma models; neuroinflammatory potential under investigation. |
*BBB: Blood-Brain Barrier
Rationale and Strategies for Combination Therapies
Monotherapy with selective STAT inhibitors may face limitations due to pathway feedback loops, compensatory STAT activation, and the complex cytokine milieu of the CNS. Combination strategies are rationalized to:
- Achieve Synergistic Inhibition: Target upstream activators (JAKs) and downstream effectors (STATs) simultaneously.
- Overcome Redundancy: Inhibit multiple STAT family members activated in parallel.
- Modulate the Immune Niche: Combine STAT inhibition with agents targeting co-stimulatory signals or cell trafficking.
- Enhance CNS Delivery: Pair STAT inhibitors with blood-brain barrier permeabilizers or nanoparticle formulations.
Table 2: Promising Combination Therapy Approaches in Neuroinflammation
| Combination Rationale | Example Drug Pairing | Proposed Mechanism in Neuroinflammation | Experimental Model Outcome |
|---|---|---|---|
| Vertical Pathway Inhibition | JAK1/2 Inhibitor (Baricitinib) + STAT3 Inhibitor (BP-1-102) | Blocks cytokine receptor signaling & downstream transcriptional activity. | In EAE, superior reduction in clinical score vs. monotherapy; reduced Th17 cell infiltration. |
| Horizontal Pathway Inhibition | STAT1 Inhibitor (Fludarabine) + STAT3 Inhibitor (C188-9) | Concurrently inhibits IFN-γ and IL-6 driven pathogenic programs in glia. | In vitro, completely abolished microglial NO release and synergistically protected neurons. |
| Multi-Modal Therapy | STAT5 Inhibitor (AC-4-130) + S1PR Modulator (Fingolimod) | Inhibits oligodendrocyte apoptosis & sequesters lymphocytes in lymph nodes. | In cuprizone model, enhanced remyelination and reduced cortical demyelination. |
| CNS Penetration Enhancement | STAT3 Inhibitor + P-glycoprotein Inhibitor (Elacridar) | Increases brain bioavailability of the STAT3 inhibitor. | Measured 3.2-fold increase in brain [compound] in murine pharmacokinetic study. |
Experimental Protocols for Key Cited Studies
Protocol 1: Evaluating STAT Inhibition in Primary Microglial Cultures
Aim: To assess the efficacy and selectivity of a STAT inhibitor on microglial activation. Materials: Primary murine microglia, LPS (100 ng/mL), IFN-γ (50 ng/mL), test inhibitor, ELISA kits (TNF-α, IL-6), Western blot reagents (p-STAT1, p-STAT3, total STATs), NO assay kit. Method:
- Isolate and culture primary microglia from P1-P3 mouse pups.
- Pre-treat cells with varying concentrations of the STAT inhibitor (e.g., 1, 5, 10 µM) or vehicle for 1 hour.
- Stimulate with LPS (for STAT3/TNF-α/IL-6) or IFN-γ (for STAT1/NO) for 6-24 hours.
- Quantification: Collect supernatant for NO (Griess assay) and cytokine (ELISA) analysis. Lyse cells for Western blot to measure phospho- and total-STAT levels.
- Analysis: Calculate IC50 for phospho-STAT inhibition and cytokine suppression. Use one-way ANOVA with Dunnett's post-test.
Protocol 2: In Vivo Efficacy in Experimental Autoimmune Encephalomyelitis (EAE)
Aim: To evaluate the therapeutic effect of a STAT inhibitor or combination therapy. Materials: C57BL/6 mice, MOG35-55 peptide, Complete Freund's Adjuvant, Pertussis toxin, test compounds, clinical scoring scale. Method:
- Induce EAE in mice by subcutaneous immunization with MOG35-55/CFA, followed by pertussis toxin injections.
- At disease onset (clinical score ≥1), randomize mice into treatment groups (n=8-10): Vehicle, Monotherapy A, Monotherapy B, Combination A+B.
- Administer compounds daily via oral gavage or intraperitoneal injection.
- Score mice daily for clinical signs (0: normal, 5: moribund) for 25-30 days post-immunization.
- Endpoint Analysis: Perform histopathology on spinal cords (H&E, LFB, IHC for CD3+ T-cells, GFAP+ astrocytes). Isolate CNS-infiltrating cells for flow cytometry (Th1/Th17 cells).
- Statistics: Compare mean daily clinical scores (two-way ANOVA) and area under the curve (one-way ANOVA).
Visualization of Pathways and Workflows
Title: JAK-STAT Pathway & Therapeutic Inhibition in Neuroinflammation
Title: Experimental Workflow for STAT Inhibitor Evaluation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for JAK-STAT Neuroinflammation Research
| Item Name | Supplier Examples | Function in Research |
|---|---|---|
| Phospho-STAT Specific Antibodies | Cell Signaling Technology, Abcam | Detect activated (tyrosine-phosphorylated) STAT1 (Y701), STAT3 (Y705), STAT5 (Y694) via Western blot or IHC. Crucial for inhibitor validation. |
| Luminex Multiplex Cytokine Panels | R&D Systems, Bio-Rad | Simultaneously quantify multiple cytokines (e.g., IL-6, TNF-α, IFN-γ, IL-10) from small volumes of cell supernatant or CSF. |
| Selective STAT Inhibitors (Tool Compounds) | MedChemExpress, Selleckchem | Pharmacological probes for in vitro and in vivo target validation (e.g., BP-1-102, Stattic, AC-4-130). |
| JAK/STAT Pathway PCR Array | Qiagen, Bio-Rad | Profiling expression of 80+ genes related to the JAK-STAT pathway for uncovering transcriptional consequences of inhibition. |
| Primary Microglial Culture Kits | ScienCell, Miltenyi Biotec | Isolate and culture highly pure primary microglia from rodent brains, essential for physiologically relevant in vitro models. |
| Blood-Brain Barrier Penetration Prediction Kit | Thermo Fisher, Corning | In vitro assays (e.g., PAMPA-BBB) to estimate compound permeability across the BBB during early development. |
| STAT DNA-Binding ELISA Kits | Active Motif | Quantify STAT protein binding to specific DNA consensus sequences, directly measuring functional inhibition of transcription. |
| CNS Penetrant Formulation Reagents | Avanti Polar Lipids, Sigma-Aldrich | Lipids and polymers for creating nanoparticle or liposome formulations to enhance brain delivery of inhibitors. |
Conclusion
The JAK-STAT pathway is a master regulator of neuroinflammation, with its activation mechanism serving as a convergent signaling node for detrimental glial responses and neuronal damage. From foundational ligand-receptor interactions to complex cross-talk, understanding this pathway is paramount. Methodological advances now enable precise dissection of its cell-specific roles, directly informing the development of blood-brain-barrier-penetrant JAK inhibitors. While troubleshooting experimental variability remains crucial, validation across models and human studies strongly supports JAK-STAT as a high-value therapeutic target. The future lies in developing CNS-optimized, cell-type selective JAK-STAT modulators and combination strategies. Success will depend on continued integration of basic mechanistic research, sophisticated disease modeling, and biomarker-driven clinical trials to translate pathway inhibition into effective neuroprotective therapies.